Great strides have been made in the lab with chemical systems that ape life’s behaviour
For Leslie Orgel, understanding the chemical processes that led to the first living organisms was one of chemistry’s most intriguing puzzles. What happened billions of years ago to transform simple inorganic chemical compounds into genetic material shared by all growing, evolving lifeforms?
As it became apparent in the 1960s that RNA played a crucial role in the transfer of genetic information, Orgel was one of the first to suggest that the earliest ‘living’ entities might have been entirely based around RNA – a theory that would later become known as the ‘RNA world hypothesis’.
The creation of a self-replicating system of ligases actually fulfils Nasa’s working definition of life
Irene Chen, University of California, Los Angeles
Writing in 1968, he proposed that RNA molecules could create new versions of themselves without the help of catalytic proteins that perform so many roles in contemporary biology. If so, it seemed plausible that this RNA could also modify the chemistry of its environment – providing a blueprint for the synthesis of polypeptides and other chemicals essential for life.
Life changing experience
The holy grail, Orgel wrote in 1995, would be to repeat this in the lab. The key would be to find a macromolecule – RNA or otherwise – capable of templating its own synthesis with enough speed and precision that it could grow exponentially. But there would still need to be scope for mutations that could be selected by a Darwinian process.
The earliest example of this unnatural chemical evolution was demonstrated by Sol Spiegelman in 1967. In a series of protein-catalysed experiments, he revealed that populations of RNA molecules could replicate and evolve and thrive under different test-tube environments.
As Orgel pondered this holy grail in the mid-1990s, the field of in vitro evolution was beginning to blossom. Meanwhile, the discovery of ribozymes had added weight to the notion of an RNA world, confirming that RNA molecules could catalyse biochemical reactions. And work on the structure of the ribosome was providing further evidence of RNA’s crucial role in protein synthesis.
‘By the time you get to 1995, there wasn’t yet the crystal structure of the ribosome – but based on Harry Noller’s work, it was pretty clear that the catalytic part of the ribosome was RNA,’ says Gerald Joyce, a pioneer of modern in vitro evolution techniques, who completed his PhD at Orgel’s Salk Institute lab. ‘Leslie called this the smoking gun – there really was an RNA world. Maybe not out of the soup, but there really was one.’
But nobody had found an RNA unit capable of unaided replication.
Copy that
That would change in 2002. Working at the Scripps Institute, Joyce and his then postdoc Natasha Paul redesigned a ribozyme to make a product that was identical to the parent. ‘The Natasha Paul paper was important because it showed that, in principle, this molecule could copy itself,’ says Joyce. The relatively simple system wasn’t capable of growing exponentially, but it proved the concept – RNA enzymes could reproduce themselves.
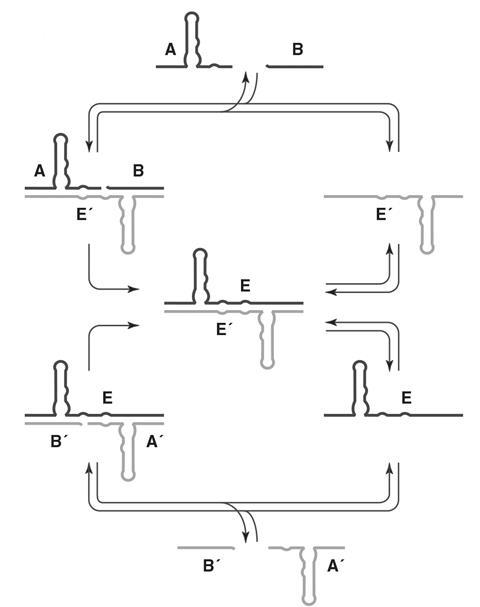
Joyce’s lab continued to develop the system, adapting the ribozyme into a cross-catalytic format in which two complementary ribozymes would promote each other’s synthesis from a pool of four substrate molecules. Initial attempts were slow and the tendency of the substrate molecules to form unreactive complexes prevented exponential growth.
The next major breakthrough came when Joyce’s PhD student, Tracy Lincoln, used in vitro evolution techniques to boost the catalytic prowess of the RNA enzymes.
‘Tracy’s paper – what people call the Lincoln replicator – was the first time we actually went exponential,’ says Joyce. ‘Furthermore, we could have populations of things going exponential simultaneously with inheritance – but very limited information content.’
‘[Lincoln and Joyce’s] creation of a self-replicating system of ligases actually fulfils Nasa’s working definition of life – “a self-sustaining chemical system capable of Darwinian evolution”,’ notes Irene Chen, whose lab at the University of California, Los Angeles, combines in vitro evolution techniques with high-throughput sequencing to map out and predict the outcomes of evolving chemical systems.
Critical success
Orgel passed away in 2007, more than a year before the Lincoln study was published. But Joyce, who had remained close friends with him, had shared the results with him as they came in from the lab. ‘He knew that it went critical, as we called it,’ says Joyce. ‘And when I showed him that, he said: “Well, that’s it then. That’s it then.” – meaning the holy grail.’
But Joyce wasn’t quite satisfied. While the Lincoln replicator could grow exponentially the next step would be to prove that it could evolve. ‘We’ve been the first to say that’s not enough – it has to have inventive capability. So it has to be somewhat open-ended, self-sustaining Darwinian evolution that is inventive – that can create new function out of out of evolutionary opportunity.’
Recent studies by Joyce’s lab have accelerated the rate at which RNA replicators amplify, and have now demonstrated ribozymes capable of making large functional molecules. The next challenge is to drive up the accuracy.
‘We’ve got enough efficiency, we’ve got regiospecificity… It makes the right linkage,’ Joyce explains. ‘It’s now quite efficient. It’s broadly sequence general – there’s some sequences it chokes on – but it’s so sequence general, it would be open-ended evolution. It’s all about fidelity. And if we can get the fidelity up from the current 97% to 99–99.5%, then it would be there.’
Order from chaos
While RNA is the prime candidate for the first genetic material, question marks still surround its emergence on the early Earth. Writing in 1999, Orgel and Joyce themselves suggested that the formation of oligonucleotides on the primitive Earth would be a ‘near miracle’.
But just three months after the Lincoln replicator results were announced there was a second major boost for proponents of the RNA world. John Sutherland’s lab at the University of Manchester showed how phosphate could catalyse a sequence of reactions and then be incorporated into RNA building blocks. ‘Leslie saw some of the earliest Sutherland work before he passed away, and really thought it was quite a breakthrough,’ says Joyce.
Six years later, Sutherland’s lab – now at the MRC Laboratory of Molecular Biology in Cambridge – would reveal how not only ribonucleotides, but also amino acids and lipids could all be generated at the same time using common chemistry. The production of lipids is particularly important in answering another big question: where might the initial replicating systems have emerged?
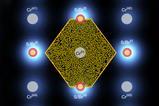
There have been suggestions that RNA molecules concentrated on mineral surfaces, porous rocks or inside emulsion droplets. Another possibility is that RNA evolution took place inside membrane-bound units. Such structures have an instinctive appeal to researchers as they resemble biological systems, and offer a clear evolutionary path to cellular life.
Holding it together
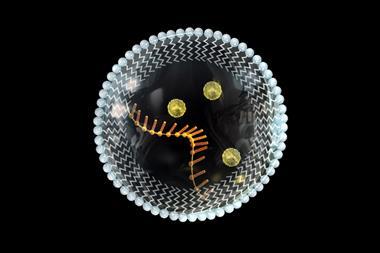
For Harvard University’s Jack Szostak, compartmentalisation is essential. Without it, RNA-catalysed products would simply drift away from the ribozyme that made them, whereas keeping them nearby supports the positive feedback required in a Darwinian process.
Over the last two decades, Szostak’s group has investigated protocells with membranes made from amphiphilic fatty acids. Importantly, these are permeable enough to allow substrate molecules to enter without requiring the complex transport mechanisms that exist in modern cells. His group has also created RNA-containing protocells that display increasingly life-like functions – including growth and division, and competing for essential chemical resources.
‘Jack’s development of a model protocell points toward a system that might someday have enough complexity to achieve open-ended evolution,’ says Chen, who previously worked in Szostak’s lab. The RNA molecules used in these experiments are relatively simple, but they are blurring the boundaries between synthetic chemistry and biology.
‘I believe that one of Jack and [Gerald’s] great conceptual contributions is to focus efforts in this area on concrete experimental models, in particular, RNA and protocells,’ says Chen. ‘The models might seem simple to a biologist, but their behaviour is surprisingly and wonderfully rich.’
Co-auth network caption: This chart shows authors of important papers in the area of origins-of-life chemistry. Each researcher is connected to other researchers they have co-authored papers with, while the colour of the node represents when their earliest highly-cited paper was published, across the period from 1995 onwards. The chart’s unusual appearance reflects the diverse topic areas that contribute to origins-of-life chemistry. The large hub in the centre of image appears to be a network of researchers whose interests focus on using astro- and geochemical data to understand the emergence of life on the early earth. John Sutherland and Jack Szostak appear as prominent nodes near the centre of the image. Leslie Orgel can be found near the top-right corner.
Cite-net caption: In this graphic researchers are linked to those other researchers who regularly cite their work. The larger the node the more often they have been cited in origins-of life chemistry papers over the last 25 years. The colour denotes when their first key paper was published, from 1995 onwards. Leslie Orgel is the largest node at the centre of the graphic, which is perhaps unsurprising given his broad interests – not only in replicating systems and informational polymers, but also in the prebiotic chemistry of the early Earth. To the right of Orgel, we can find Stanley MIller, known for his work on the Miller–Urey experiment showing how organic compounds could be synthesised from inorganic precursors under prebiotic conditions. Joyce and Szostak appear as prominent nodes to the left of Orgel, while the group to the far-left of the image is dominated by researchers known for their work on systems chemistry.
Geo caption: From this map of author locations for key origins-of-life chemistry papers, it is clear to see that most of the interest in this topic is centred in North America and Europe. In comparison to many of the other holy grail subject areas, there appears to be relatively little research activity in China for this field.
Citation race caption: This chart shows the top papers from the last 25 years that relate to the emergence of replicating chemical systems. The most highly cited paper is a review written by Vincenzo Balzani, Marcos Gómez-López, and Fraser Stoddart supramolecular assemblies and molecular machines. The next paper is another review on prebiotic chemistry and RNA’s origins by Leslie Orgel, the author of the original holy grail paper. Tracey Lincoln and Gerald Joyce’s landmark paper on ‘the Lincoln replicator’ appears in sixth place, underlining its huge importance to this grail area.
Additional information
For an explanation of how these data visualisations were built, read Behind the data
Further reading
1 L Gabrielli, D Núñez-Villanueva and C A Hunter, Two-component assembly of recognition-encoded oligomers that form stable H-bonded duplexes, Chem. Sci., 2020, 11, 561 (DOI: 10.1039/c9sc04250d)
2 L Gabrielli and C A Hunter, Supramolecular catalysis by recognition-encoded oligomers: discovery of a synthetic imine polymerase, Chem. Sci., 2020, 11, 7408 (DOI: 10.1039/d0sc02234a)
3 G Creusen et al, Scalable one-pot-liquid-phase oligonucleotide synthesis for model network hydrogels, J. Am. Chem. Soc., 2020, DOI: 10.1021/jacs.0c05488
4 C A Figg et al, Controlling the DNA hybridization chain reaction, J. Am. Chem. Soc., 2020, 142, 8596 (DOI: 10.1021/jacs.0c02892)
5 E A J Post and S P Fletcher, Dissipative self-assembly, competition and inhibition in a self-reproducing protocell model, Chem. Sci., 2020, 11, 9434 (DOI: 10.1039/d0sc02768e)
6 Y Lyu et al, Protocells programmed through artificial reaction networks, Chem. Sci., 2020, 11, 631 (DOI: 10.1039/c9sc05043d)
7 M A Cardona and L J Prins, ATP-fuelled self-assembly to regulate chemical reactivity in the time domain, Chem. Sci., 2020, 11, 1518 (DOI: 10.1039/c9sc05188k)
8 E C Fischer et al, New codons for efficient production of unnatural proteins in a semisynthetic organism, Nat Chem Biol, 2020, 16, 570 (DOI: 10.1038/s41589-020-0507-z)
9 G Houlihan et al, Discovery and evolution of RNA and XNA reverse transcriptase function and fidelity, Nat. Chem., 2020, 12, 683 (DOI: 10.1038/s41557-020-0502-8)
10 W Ding et al, Chimeric design of pyrrolysyl-tRNA synthetase/tRNA pairs and canonical synthetase/tRNA pairs for genetic code expansion, Nat. Commun., 2020, 11, 3154 (DOI: 10.1038/s41467-020-16898-y)

Searching for the holy grails of chemistry

How has science progressed over 25 years?
- 1
- 2
- 3
 Currently
reading
Currently
reading
Artificial replicators that mimic life offer new insights into its origins
- 5
- 6
- 7
- 8
- 9


































1 Reader's comment