In a chance discovery, researchers have for the first time observed the emergence of something akin to metabolism in molecules that also self-replicate.1 The replicator molecules catalyse a reaction that produces compounds that help to make the replicator’s own building blocks. It might be one of the most lifelike chemical systems ever created, combining two of life’s three essential features – replication, metabolism and compartmentalisation.
‘What it can teach us is that those functions that we see in living systems, they are not unique to life as we know it,’ says origin of life scientist Kamila Muchowska from the University of Strasbourg, France, who wasn’t involved in the work.
Complete coincidence
Although self-replication is usually associated with DNA, the behaviour has been seen in very different looking chemical systems – for example in rotaxanes. This raises the intriguing possibility of creating completely synthetic lifeforms that tick all three boxes for life: replication, the use and storage of energy to perform energetically unfavourable, or endergonic, reactions, and keeping all of these functions contained to protect them from parasitic lifeforms.
Systems that either replicate or perform metabolism-like reactions have existed before, and some can even switch from one function to the other upon addition of chemicals or environmental changes. But a team around Sijbren Otto from the University of Groningen in the Netherlands has now created molecules that do both at the same time.
The system’s basic building block is a benzene-1,3-dithiol attached to a short peptide chain. In the presence of oxygen, the thiols oxidise and form disulfide macrocycles of different sizes, mostly trimers and tetramers. The rings constantly exchange thiol building blocks between each other, so there is a small chance of larger rings forming, in particular six-membered ones. These hexameric rings assemble into stacks or fibres that keep growing at both ends – this is the replicator. Upon mechanical agitation, the longest fibres break into two and then start growing again individually. As more rings are incorporated into stacks, the solution equilibrium replenishes ones of that size. What follows is exponential self-replication.
But Otto and his colleagues have now discovered – by coincidence – that the replicator stacks can catalyse reactions that the building blocks cannot. On the stacks’ surface, positively charged lysine side chains sit in close proximity, which enhances their Brønsted basicity.
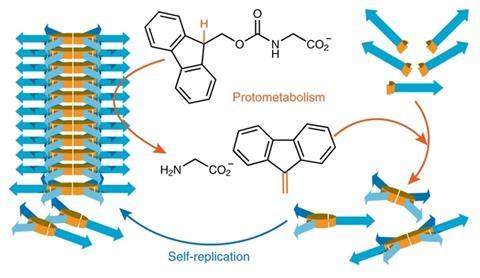
In a Brønsted base-catalysed reaction, the replicators cleave a fluorenylmethoxycarbonyl (Fmoc) protected glycine, creating dibenzofulvene. This compound speeds up the dithiol building block’s oxidation and hence the replicator’s formation. ‘We now have replicators that are able to catalyse reactions that produce products that help with making molecules from which the replicator grows – that’s protometabolism,’ Otto says.
‘It’s a chance discovery, the system doing something that was never designed in,’ says Otto. ‘We’ve been working on these molecules for 10 years, we could have discovered this 10 years ago. We weren’t looking the right place, but it was there.’
Life as we don’t know it
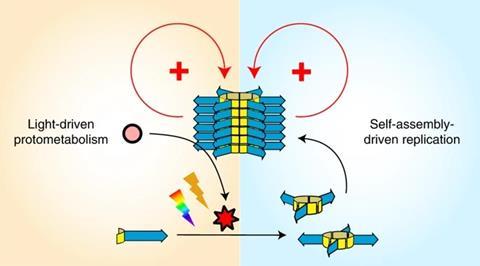
In a separate study,2 Otto’s team offered the replicator a photosensitiser – rose bengal or tetraphenylporphyrin – instead of Fmoc–glycine. The replicator fibres bind the sensitiser, giving it the ability to photocatalytically produce singlet oxygen, which in turn promotes oxidation of its building blocks.
Muchowska is hesitant to call the system protometabolic – it is missing the ability to store energy and perform endergonic processes – but says that ‘the fact that they’re recruiting a cofactor to perform catalysis within the self-replicating system is really important, I think, to start looking at the emergence of biocatalysis in a living system’.
While Otto’s system is not trying to imitate biology, it might help researchers solve a persistent puzzle surrounding life’s origin on Earth. Eigen’s paradox states that accurate replication needs complex machinery, but to make complex replicators through evolution requires accurate replication – a chicken-and-egg problem. ‘There’s no real evidence yet of experimental systems that solve the Eigen paradox,’ says Otto. ‘But we’d love to try with our system.’
One idea to solve the paradox is to not store all the information in one long molecule, but rather in a cooperative systems of multiple molecules each carrying a small amount of information, explains Yu Liu who models complex chemical systems at the University of Glasgow, UK. The community becomes a unit of selection, rather than the individual.
But building these systems could also help researchers recognise alien life, were they ever to come across it on other planets, Liu says. And life’s mechanisms can be abstracted even further to mechanical lifeforms or those purely contained within a computer, akin to John Conway’s Game of Life.
‘How far is [our system] away from being alive?’ wonders Otto. ‘I think we need to integrate two more things: compartmentalisation and open-ended evolution. We’re now really probing questions that relate to how we can evolve the system if we offer structural variation possibilities.’
References
1 J Ottelé et al, Nat. Catal., 2020, DOI: 10.1038/s41929-020-0463-8
2 G Monreal et al, Nat. Chem., 2020, DOI: 10.1038/s41557-020-0494-4
















No comments yet