Molecular computing systems could improve disease diagnosis – and even hack living cells, finds Andy Extance
Credited with coming up with the term ‘computer virus’, by the early 1990s US computer scientist Len Adleman had moved his attention to one of their most insidious biological namesakes: HIV. The University of Southern California researcher had already co-invented an algorithm in the 1970s that still powers safe online transactions and had even consulted for the 1992 hacker movie Sneakers. But for all the reputation he’d earned, no-one would listen to his ideas on HIV.
To ‘learn the language of real HIV researchers better’ Adleman spent a summer in his university’s molecular biology lab. The outcome was a new field of science that distracted him from HIV completely. ‘I see that DNA is a mathematical object – it’s just a string of characters,’ Adleman recalls. ‘I see techniques like the polymerase chain reaction as computations.’
When his early experiments showed DNA could compute and might even be used to tackle hard maths problems – including breaking cryptographic schemes – better than silicon, Adleman set up his own lab focusing on DNA computing. Though his explorations earned him the mantle ‘father of DNA computing’, after a decade he concluded that DNA would not replace silicon for solving hard maths problems. And after his final paper on the topic in 2002, Adleman withdrew from active research in the adolescent field.
Abandoned by its father, its limitations exposed, DNA computing seemed to be in crisis. But thanks to foundations laid partly in Adleman’s lab, DNA computing and molecular programming now mean humans could rationally design controls for biochemical systems – even those inside living organisms.
Computing, but not as we know it
DNA computing appeared explosively in 1994, when Adleman showed DNA strands could solve the ‘travelling salesman’ problem.1 The challenge is finding a route between various cities, passing through each only once. Because the number of possible answers increases exponentially with each extra city, the problem rapidly eats up conventional computers’ resources.
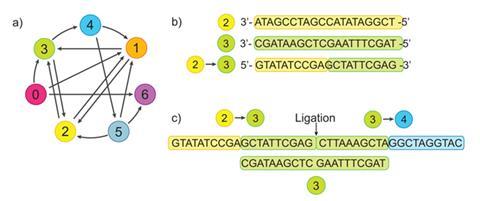
Adleman used DNA strands with 19 different sequences, each 20 nucleotide bases long, to represent a map. Seven of the sequences represented cities and 12 sequences represented possible paths between them. Not every city was connected to every other. The first 10 nucleotides of each path strand are complementary to the last 10 nucleotides of the sequence representing the first city that this path connects. The last 10 nucleotides of a path strand are complementary to the first 10 nucleotides of the sequence representing a second city that the path connects. Therefore each path could pair up into a DNA double strand with two cities and each city other than the first and last would likewise pair up with two paths. And thanks to the presence of a DNA ligase enzyme, each stage could be joined together into ‘journey’ sequences answering the problem.
Over seven days of lab work Adleman mixed the different cities and paths, getting a selection of assembled journeys. He used enzyme-driven polymerase chain reaction (PCR) amplification to create more copies of routes linking the desired starting and ending cities and purified out any that didn’t visit each city once. Amplifying the few remaining strands by PCR again gave enough DNA that the answers could then be read off as bands on a gel chromatograph.
In these 50 picomolar scale reactions around 3 × 1013 copies of each starting oligonucleotide were available to do computation in parallel. If each binding were considered to be a computational operation, during the week-long experiment around 4 × 1014 operations were performed, each using a tiny amount of energy. At the time, the best silicon computers could do around 1012 operations per second, or around 7 × 1014 in two hours. With further improvements seeming possible, the media and scientists alike bubbled with excitement.
Yet scale-up proved tricky. In 2002, Adleman and his team published their largest DNA calculation, which involved over a million different sequences.2 At the time, they wrote that ‘this problem may be the largest yet solved by non-electronic means. Problems of this size appear to be beyond the normal range of unaided human computation.’
However, University of Texas at Austin, US, biochemist Andy Ellington later underlined that, if those unaided people got help from electronic computers, the problem could be ‘trivially solved’.3 A stern critic of parallel DNA computation, Ellington once called it ‘a good example of a failed technology arc’. Its downfall came from unwanted interactions. Every time wrong pairings form, the system swaps information for nonsense. When you’re hunting one ‘right answer’ molecule in a million that can be fatal. ‘It quickly became apparent that it would require truly extraordinary achievements in molecular engineering to have DNA act with the fidelity that your average handheld calculator accessed,’ says Ellington.
Attempting the extraordinary
By then Erik Winfree was collaborating with Adleman, while pursuing his PhD on programming the self-assembly of structures like DNA tiles at the California Institute of Technology (Caltech) in Pasadena, US, where he now has his own team. That experience showed him how DNA could perform an initially more modest, and perhaps even less familiar, type of computation. From there, his Caltech team would develop chemical reaction networks where DNA strands switch partners, exploiting an existing concept known as ‘toehold-mediated strand displacement’ to produce circuits.
The toehold is a sequence of nucleotides dangling off the longer strand of two bound, uneven-length oligonucleotides. If an incoming oligonucleotide has the right sequence to bind with the longer partner, including the extra dangling ‘toehold’, it can rapidly displace the shorter partner. Using this enzyme-free, non-covalent approach, in 2006 Winfree’s team published details of AND, OR and NOT circuits – the most basic logical elements.4
An AND gate produces an output signal when two input signals are both turned on. The Caltech team’s version comprises three parts, two shorter strands bound to a longer one. A first oligonucleotide input removes one of the short strands through toehold displacement. That opens up a toehold on the remaining pair of strands, one of which is fluorescent and the other fluorescence-quenching. The second input oligonucleotide separates these, ultimately causing the fluorescence that is the circuit’s readout.
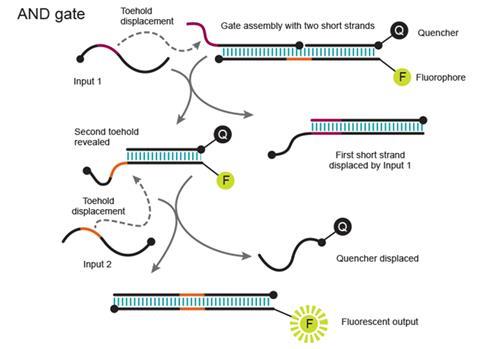
This helps minimise unwanted ‘crosstalk’, caused by the incorrect DNA pairings that plagued Adleman’s approach, Winfree observes. He underlines that chemical systems are not like electronics, where signals are carried by electrons on separate wires that cannot interfere. Instead, signals take the form of molecules floating around in solution, bumping together randomly. Carrying the signals into an AND gate as sequences only able to do one specific displacement comes close to the separation achieved by wires.
However, in strand displacement, making DNA behave digitally doesn’t stop there. ‘If you have a faulty signal that should be a high concentration – a digital ‘one’ – but it’s only a middle-high concentration, you need to amplify it,’ explains Winfree. ‘When the problem is an unwanted signal, one way to isolate the desired signal is to have a small amount of competitor that will consume the unwanted one. The desired signal can then propagate through the circuit. After the correct signal gets past the competitor, you want a catalytic reaction that will amplify what’s left up to a standard value.’
By 2011, Winfree and co-worker Lulu Qian had produced a circuit where fluorescence reported square roots of numbers up to 15, rounded down to the nearest integer.5 The circuit brought together several strand displacement logic gates, each comprising a few oligonucleotides, with the overall circuit containing 130 different strands. Winfree and Qian read off the results in binary from the colours produced by four different fluorescent outputs. ‘At every AND or OR gate there’s a competition process followed by catalytic amplification,’ Winfree underlines.
Soup-er computer
At Microsoft Research in Cambridge, UK, Andrew Phillips and his colleagues have built a programming language called DSD for designing ‘DNA strand displacement’ systems like Winfree’s team’s. It integrates with an existing approach known as Chemical Reaction Networks (CRNs) that models how molecular components interact. Phillips compares this to programming on a PC, only embodying instructions in DNA rather than on a CD or hard drive.

‘On a PC you write a program in a language such as C, then compile it down to some intermediate language, then compile that down to zeroes and ones that a computer can execute. DNA is like zeroes and ones, CRNs as the intermediate language, and on top of that is DSD. That allows us to come up with strand displacement algorithms, compile CRNs and then build those CRNs out of molecules. Designing the algorithm is the challenge. A typical computer program is like a sequential recipe – a program in chemistry is more like a soup.’
In 2013, the Microsoft Research team used DSD with two US-based academics: David Soloveichik at the University of California, San Francisco, and Georg Seelig at the University of Washington in Seattle. Together the teams designed and built strand displacement control module circuits from scratch that could work with DNA sensors and motors.6 To show what computing can be done with strand displacement, which Soloveichik and Seelig helped invent years earlier as part of Winfree’s Caltech team, one controller implemented an ‘approximate majority’ algorithm.
This algorithm reacts two populations of possible output strands together so they can ‘agree on which population is larger’ by forming intermediates that can amplify either output. This arrangement consumes both signal strands, effectively cancelling them against each other, eventually leading to the one that’s present in higher concentration becoming completely dominant. The same algorithm is used in computing across networked systems, and in cells when they divide, Phillips notes, though in DNA programming it’s still slow. ‘This algorithm takes 15 hours,’ he admits. ‘You wouldn’t have a phone that runs on these algorithms. Their advantage is that they can potentially run inside cells.’
Making DNA better?
Rather than chemically synthesising oligonucleotides, Soloveichik and Seelig made their circuits by inserting DNA into Escherichia coli’s genome, breeding the bacteria, and then extracting their DNA again. That overcomes another source of information loss arising from incorrectly synthesised sequences, Phillips says, at comparable cost.
Winfree calls this ‘very impressive and important’, especially for ‘analogue’ computations that allow values between the ones and zeroes specified by digital computing. But while bacteria produce better-functioning circuits, errors in oligonucleotides made chemically ‘are not deal-breakers yet’. ‘In my lab, we design a sequence, send an email to a company and a few days later they send us the DNA. Our typical experiments use maybe $100 (£66) worth of reagents. We don’t really care how they made it, we care that it doesn’t mess up experiments. But I don’t think that synthesis is a major roadblock.’
In fact, Winfree doesn’t see any insurmountable roadblocks to progress along the path to making more complex molecular computing machines using DNA. His collaborators have already designed and built a biochemical oscillator ‘clock’ from DNA, RNA and enzymes and placed it in oil droplets that acted as simple ‘artificial cells’.7 This is a step towards ‘molecular robots,’ Winfree explains. ‘We want a device as small as a cell that can navigate through a chemical environment, emit chemicals that interact with its environment and have chemical sensors to see what’s in it.’ Using DNA computing to deliver the control needed to get the system to do the right thing at the right time is molecular programming, he says.

Progress in programming DNA systems that can interact directly with biochemicals has even converted Ellington. ‘We’re doing a very simplistic thing, to filter noise in molecular diagnostics,’ he says.8 ‘One problem is that if you amplify a target strand, you also amplify unwanted ones. How do you filter signal from noise?’ One answer is circuits that do basic DNA computing using strand displacement. In filtering out such noise, the approach’s selectivity performs a similar function to preventing crosstalk between different signals in Winfree’s circuits.
Ellington is now working with Paratus Diagnostics, which is also based in Austin, to commercialise this technology in devices that can diagnose illnesses from RNA or small molecules in saliva or urine. Ellington hopes it could enable ‘home doctor’ products – but is characteristically realistic on the chances of success. ‘I have no fear the marketplace will tell me exactly what it thinks in the not-distant future,’ he laughs.
Yet DNA programming is poorly suited to some possible applications, he adds. ‘I draw the line at bulky DNA drug delivery devices – they are completely untenable. They cost too much – the field is up against small molecules made for pennies that have great therapeutic efficacy.’
Life-changing results
Peng Yin from Harvard Medical School’s Wyss Institute in Boston, US, also sees diagnostics as important – but has a much grander long-term vision. ‘You could envision a nanodevice that, upon detecting a disease signature, such as proteins or RNA molecules, turns on the production of proteins that could diagnose it by changing the diseased cell’s colour, or cure it by killing the cell,’ he says. In October 2014 Yin’s team drew closer to this scenario, publishing RNA circuits designed from scratch that control protein synthesis in an E. coli cell.9

The scientists used strand displacement both to provide the low crosstalk levels needed to make circuits function in cells, and create a powerful design framework. That helped when peer reviewers asked for a daunting number of extra experiments. ‘We said, “Our systems are already optimised, so we might be able to do this,”’ Yin recalls. ‘The postdoc leading the work, Alex Green, spent two and a half years to set up the basic principles. He finished the extra experiments in three months.’ Green and his colleagues were also able to translate this knowledge to make a paper-based Ebola sensor in ‘just a couple of days’.10 ‘That shows remarkable programmability and robustness, and that’s exciting for synthetic biology,’ Yin enthuses. And while the paper sensors are still at an early stage, a number of biotech companies are interested in them, he reveals.
Yin’s team engineered E. coli to produce two RNA components itself. One is a ‘toehold switch’ RNA loop, containing instructions for protein manufacturing that opens up in the presence of a second ‘trigger’ component. Once the loop is open a ribosome can bind, read it, and make the protein it codes for. Though this is simple if/then logic – if detecting RNA A then make protein B – Yin underlines its power. Previous RNA systems adapted from nature already perform similar functions, but usually with a relatively narrow 50-fold difference between protein concentrations in the on and off states. On average, in the new system the difference is 400-fold.
Yin underlines that this demonstrates the power of rational design. ‘It is quite amazing that these synthetic gene regulators way outperform those adapted from regulators optimised by evolution,’ he enthuses. ‘Evolution works, but physics rules. This is the first substantial demonstration that the principles that we’ve developed for molecular programming in a test tube can be applied to a living cell. But this is just a starting point. We have ongoing work to demonstrate more sophisticated in vivo computing than ever before.’
In computing terms, DNA and RNA’s capabilities are still primitive. But molecular programming’s power to interface with biological systems is already clear, in disease testing, creating molecular robots and even modifying living cells. Inspired by the potential to hack secure electronic communications between humans, DNA computing and molecular programming are now helping hack into perhaps the greatest mystery of all – life.
Andy Extance (@andyextance) is a science writer based in Exeter, UK













No comments yet