The UK pioneered a forensic process to identify suspects from tiny amounts of DNA, but occasional flaws had big consequences. Andy Extance pieces together the whole story for the first time
In 2002, 10 years after the notorious unsolved murder of Rachel Nickell, Andrew McDonald entered a small briefing room in Abingdon, UK. There, in the offices of Forensic Alliance, Metropolitan Police Detective Chief Inspector Richard Brooks shared details about how the 23-year-old was killed while walking with her two-year-old son and dog on Wimbledon Common in London on July 15, 1992. She was sexually assaulted and died with 49 stab wounds.
After Brooks had spoken, a reporting scientist from the UK Forensic Science Service (FSS) London laboratory handed a file to McDonald, DNA lead at Cellmark Forensic Services in Abingdon. ‘I distinctly remember him saying “Good luck, you won’t find anything”,’ McDonald tells Chemistry World.
McDonald was part of a Forensic Alliance team assembled by company founder Angela Gallop, who has a strong record of using forensics to resolve cold cases. They became involved long after the Metropolitan Police had focussed on a local man called Colin Stagg, who they arrested and charged but who was acquitted in September 1994.
After receiving the first samples in 2003, McDonald noticed that a key DNA test registered no DNA at all – not even Nickell’s. ‘We just thought, well, that’s absolutely mad, because this was taken directly from Rachel’s skin,’ recalls Gallop. ‘There should have been at least lots of her own DNA there.’
When we got the results back, we had an indication of something at a much lower level from another individual
McDonald was looking at results from a DNA profiling method known as a low copy number (LCN) test. The FSS had started devising LCN testing in 1998 to deal with cases where there was very little DNA material available, then applied it to the Nickell case. DNA profiling relies on the polymerase chain reaction (PCR) method, best known to the public today for helping detect Covid-19. PCR copies DNA in forensic samples, boosting it to measurable levels. LCN tests push this amplification to its limits, and McDonald had seen that this sometimes stopped the test from working properly.
In the test sample which returned no result, there was much more material than expected. The FSS scientists had not realised this because, to avoid using more than needed, the FSS scientists had not measured the quantity. With so much sample, it was far likelier to contain substances that could inhibit the polymerase enzyme central to the PCR method and stop it working.
McDonald re-ran the same test at Cellmark, but using a smaller sample, thinking nothing would happen. ‘When we got the results back, we had now a really strong profile matching Rachel Nickell – with an indication of something at a much lower level from another individual, possibly a male individual,’ McDonald says.
That finding was pivotal for the Rachel Nickell case and for DNA profiling in general. The FSS reported potential issues, after which the UK’s forensic science regulator commissioned a review that highlighted when LCN worked reliably and when it didn’t. The review’s findings then triggered the FSS to secretly revisit 2180 affected cases. Meanwhile, a case involving a terrorist bombing in Northern Ireland highlighted different issues with the LCN method. This is the first account to bring these events together and show how they drove forensic scientists to refine their techniques. ‘Things have moved on enormously,’ McDonald observes.
Taking DNA to court
LCN was the world’s first forensic method for DNA profiling from a very low number of copies of genetic material. Such methods start by extracting DNA from an evidence sample, explains John Butler, research scientist and DNA forensics expert at the National Institute of Standards and Technology (NIST) in Maryland, US. That might involve washing the sample in solvents like phenol, chloroform, or methanol, though in the 1990s UK forensic scientists often extracted DNA with a resin called Chelex 100. After extraction, scientists use many rounds of PCR to amplify DNA at very specific locations, known as short tandem repeat markers, or STRs, Butler adds.
STRs are hyper-variable sequences in our genomes ‘that differentiate you and me, and we believe just about everybody we’ve ever sampled’, explains Adrian Linacre, from Flinders University in Adelaide, Australia. ‘Every one of your cells has essentially the same DNA profile. The regions we look at have no effect on what you look like. They are non-coding bits of DNA that happen to show a repetitive nature that have a four base pair repeat, something like AATG, AATG, AATG. And that repetitive nature is very useful for us.’
There are half a million repeated sequences, Linacre adds, but most of them just repeat two bases. The very first DNA profiling techniques used just four locations with four-base STRs, with LCN originally looking at 10 STR locations. ‘The more [locations] we look at the less likely it is that two people by chance will share that DNA,’ says Linacre.
Using primer molecules PCR can target these locations, with each PCR cycle doubling the amount of STR DNA. The most-used DNA profiling method in the UK in the early 2000s, called SGM Plus, used 28–30 cycles of PCR doubling. Scientists can then separate STRs by electrophoresis, labelling each with coloured fluorescent molecules. An automated analyser can then record the intensity of fluorescence, showing each STR as a peak on a software plot.
It becomes so sensitive that it will actually detect a single copy of DNA
Scientists can then relate peaks to a reference mixture run through electrophoresis alongside the sample known as an ‘allelic ladder’, explains Butler. The name refers to the fact that we normally have two copies of each gene, one from each parent, known as alleles. While each copy does the same job, they can have different makeups. The allelic ladder includes the entire known range of repeated genetic sequences. ‘That then says, for example, there’s an 11–14 genotype, so 11 repeats may be from the mother, 14 from the father,’ explains Butler. ‘You compare that to the known samples from an individual.’ That provides a likelihood ratio or random match probability, calculating the chance that they originated from the same individual.
Building on such techniques, an FSS team led by Peter Gill published its first paper on the LCN technique in 2000. Previously, the lower limit on the amount of DNA it was possible to profile with had been around 250 picograms. The FSS team showed that LCN could identify people using less than 100pg of DNA, by amplifying it with 34 PCR cycles. ‘It becomes so sensitive that it will actually detect a single copy of DNA,’ says Gill, who now leads the forensic genetics research group at Oslo University Hospital in Norway.
Getting the suspect on tape
From 2003 onwards, McDonald, Gallop and their colleagues gradually worked on the evidence from the Rachel Nickell case. But after discovering the important clue of male DNA in the original extract, they were running out of material. So, they revisited other forensic evidence, specifically ‘tapings’ from around Rachel Nickell’s vagina. Examiners had attached sticky tape, removed it again to collect debris, and stored it so the sticky side was protected, McDonald recalls. He and his colleagues were then able to swab the tape again, extracting enough DNA to repeat their earlier feat with standard SGM Plus 28-cycle profiling. They found Nickell’s DNA and around five STR sequences that didn’t match her, but this was not enough to identify a suspect.
Comparing the 12 STR sequences against the police’s ‘persons of interest’ list gave only one match
To improve on this, McDonald did trial experiments on other samples, changing the electrophoresis technique to make it 10 times more sensitive. Purifying the post-PCR mixture to remove unwanted salts and enzymes gave a total 15-fold increase in peak height. Applying this to the product of PCR from Nickell’s body taping using a profiling technique covering more STR locations identified 12 sequences from a different person. ‘That gave us enough to make a meaningful comparison between two people,’ McDonald says.
Comparing the 12 STR sequences against the police’s ‘persons of interest’ list gave only one match. That was Robert Napper, who by 2002 was held in the high security Broadmoor Hospital, after being convicted of stabbing 27-year-old Samantha Bisset to death and smothering her daughter Jazmine in their home in 1993. The match to Napper’s DNA sample was already strong, but not complete. So, McDonald and his colleagues used a profiling technique that exclusively looked at STRs on the Y chromosome, which only males have. Nickell’s DNA couldn’t interfere with this sample, and the Cellmark/Forensic Alliance team got a nearly full match with Napper’s DNA.
That prompted the team to look for more evidence among Napper’s possessions linking him to the Nickell murder, Gallop says. ‘One of the shoes that Robert Napper had could have left the footwear mark at the scene,’ she notes. ‘The paint flakes that we found in the hair combings from Rachel’s son could have come from the toolbox that he kept his tools in.’ Gallop also reveals that near the start of the investigation, a key member of her team reported to the police similarities with the Green Chain rapes. This series of 70 brutal attacks in London ended the same year that Napper went into Broadmoor, and he admitted to some of them in 1995.
LCN on trial
Together the evidence was overwhelming, convincing the police to charge Napper, who changed his plea from not guilty to guilty during the trial. In December 2008, he was convicted of Nickell’s manslaughter and sentenced to indefinite incarceration in Broadmoor. Yet the fact that LCN had missed Napper’s DNA in the first place was worrying. ‘We had demonstrated that there was a flaw with the FSS LCN technique as it was being employed at the time,’ McDonald explains. ‘Not quantifying the sample was missing potentially critical results. Alarm bells were ringing for good reason in the criminal justice system.’
In 2006 the FSS told the police and Home Office that LCN could ‘fail to identify a DNA profile from genetic material in a sample that should reveal a profile or mixture of profiles’. That helped spur the Home Office to appoint a forensic regulator to oversee forensic provision in England and Wales, recalls Linacre. The first regulator commissioned a review into the LCN technique led by Brian Caddy from the University of Strathclyde, where Linacre also worked at the time. Caddy himself was not a forensic geneticist, so he recruited Linacre, who had previously used LCN in criminal case work and Graham Taylor, head of genomic services at Cancer Research UK.
In late 2006 and early 2007 the FSS also started a secret ‘backtracking operation’ for samples that had produced no result, known as Operation Cube. Huw Turk, currently scientific lead at Cellmark, was one of three FSS scientists that went through the affected LCN tests, reported as over 5000 samples affecting 2180 cases by the Evening Standard. ‘We went back to those samples, and re-ran them at diluted concentration, because that often got around this problem of inhibition,’ says Turk. ‘It took the three of us the best part of the year to complete. A large majority still gave no result.’ Their efforts produced 26 partial or full DNA profiles of suspects in cases of murder, rape or both, according to the Evening Standard. Turk calls this ‘a very small proportion’ of the cases looked at.

Then, in December 2007, LCN issues entered headline news stories. Sean Hoey was being tried on 58 charges including the murders of 29 people in the 1998 Omagh bombing in Northern Ireland. The trial was conducted by a judge sitting in the case without a jury, due to fears that paramilitary organisations might intimidate jurors. Forensic scientists working on the case used LCN to produce a profile of Hoey, making the method a focus for the defence team. The judge, Mr Justice Weir, had therefore called the technique’s inventor, Peter Gill, to give evidence in Belfast crown court in January 2007. ‘It was the worst experience I’ve ever had,’ Gill recalls.
It might have been enough that the defence in the case put up a scientist of whom a judge in a different case wrote, ‘it is impossible to understand how [the defence expert] had sufficient expertise to be able to give evidence’. Yet the judge also questioned Gill proactively and aggressively, sometimes shouting at him. ‘I was really unable at that trial to give a proper representation of what the evidence meant and what it did not mean,’ Gill recalls.
Dropping in and out
Rather than inhibition, the case centred on two important issues for a technique as sensitive as LCN, both of which Gill and colleagues had devised a mathematical solution to. The first is known as allelic drop-out. It arises because, with so little starting DNA, the two copies of an allele might not be amplified equally. The primer DNA strands used to tell the polymerase enzyme which sequences to amplify might not bind their target. If the primer doesn’t bind at one STR location, it would look like the person from whom a DNA sample originates received the same genotype from both parents. In this case, one of their alleles has dropped out.
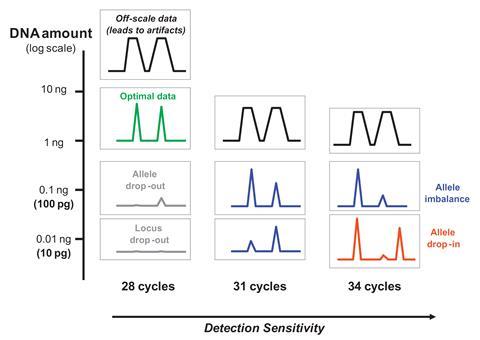
‘If an allele disappears, it obviously reduces the strength of the evidence, and you have to factor that into your calculation,’ Gill recalls. Gill’s mathematical analysis could provide a probability that the sample was really from a suspect. However, he admits that it was ‘really too complicated’ for forensic scientists to use on a day-to-day basis. ‘So, instead we use a method that required duplication,’ he says. LCN analyses were repeated, and STR alleles confirmed if they appear at least twice. ‘You can analyse simple cases using spreadsheets,’ Gill notes.
‘If you’re asked in court “Is this reproducible?” the answer would need to be “no, it’s not”,’ Linacre stresses. ‘When Justice Weir did ask Peter Gill about this reproducibility issue, he was quite honest and said “It’s not reproducible”. And immediately the judge did not like that type of concept.’
The evidence wasn’t being collected in the way that it should have been
‘That’s probably the part where he started shouting at me,’ Gill says. ‘We don’t deal with the definitive. In fact, no scientist does. But the judge did not understand that. He wanted definitive answers.’
Another issue in the Omagh bombing case was better founded, because the technique is so sensitive it easily picks up contamination. That could be one or two extra alleles from highly degraded DNA from dust, for example, known as allelic drop-in. Or it might be contamination with multiple alleles from an uninvolved person. With LCN only just beginning to be developed when the Omagh bombing happened, the forensic scientists and police involved hadn’t considered this. They had stored evidence together in bags with holes in. ‘I remember being asked, “Is it possible that DNA can go from one item of evidence to another?”,’ Gill says. ‘To which my answer was yes. The evidence wasn’t being collected in the way that it should have been.’
So, in his final December 2007 verdict, Mr Justice Weir ultimately cleared Hoey of all charges and said that he was not satisfied that the LCN method was well enough validated. Shortly following that judgment, the police suspended use of the LCN method, pending results of the Caddy review.
After four months of interviews, in April 2008, Caddy, Linacre and Taylor made 21 recommendations on the use of LCN in forensics. The headline conclusion was that it was a safe method to use. ‘In cases where you could not otherwise generate a DNA profile using 28 cycles under current technology, but you could under 34 cycles, then the process is fit for purpose and should go ahead,’ is Linacre’s summary.
Lessons learned?
Other recommendations addressed issues seen in the Nickell and Hoey cases. Concerning the Nickell case, one stated that the amount of DNA extracted should always be quantified, and technical standards for extraction and dilution agreed by all forensic science providers. Concerning the Hoey case, the review pressed for ‘national agreement’ on how to interpret LCN tests, to meet evidence standards that judges like Mr Justice Weir expected. Another recommendation focussed on minimising contamination, including a national standard for ‘DNA clean’ crime scene recovery kits, and sterile consumables in forensic labs.
Gallop highlights that such problems went beyond the UK. In Germany, DNA profiling consistently identified the same woman as a suspect in cases nationwide, who turned out to be a worker in a factory making cotton swabs. In general, Gallop adds, the Caddy review ‘really was very helpful’ in stressing the need for proper forensics training for people working on cases and for test validation and standardisation.
For Turk, the most important lesson concerning inhibition in the Nickell case was the need to quantify the DNA. ‘From that point onwards, we would have a much better idea if there was any DNA present [and] if it was at risk of inhibition,’ he explains.
A laboratory won’t spend a lot of time trying to work out why a sample hasn’t worked
Technology has also progressed rapidly since the Caddy review to mitigate other problems with profiling low amounts of DNA, adds NIST’s Butler. ‘The biggest change has been the development of probabilistic genotyping software to assign the chances of there being allele drop-out or allele drop-in information,’ he says.
In Oslo, Gill continues to work in this area, developing software packages to help implement mathematical concepts he has introduced. He warns that inhibition is always possible, however, and still happens. He also notes that people still don’t always quantify their DNA samples because of how tests are funded, especially in Europe. Tests still might not work thanks to inhibition, but given their financial constraints, forensic scientists might not go back to look at why. ‘A laboratory won’t spend a lot of time trying to work out why a sample hasn’t worked, unless it’s a really important case,’ he says.
For Gallop, the oversight made with the original LCN sample in the Nickell case ‘should never have happened’. Yet it still could, albeit for different reasons. She worries about current financial pressures that force forensic scientists to process samples too quickly. ‘You’ve actually got to think about what you’re doing and what your results mean. Forensic science is not just about testing. It’s much more complicated than that. And the consequences for getting your testing wrong are absolutely dire.’
Andy Extance is a science writer based in Exeter, UK













No comments yet