Lab-on-a-chip technology is finally seeing widespread use in analysis and synthesis. Jon Evans catches up with the progress of microfluidics research
Lab-on-a-chip technology is finally seeing widespread use in analysis and synthesis. Jon Evans catches up with the progress of microfluidics research
They’ve long been anticipated, but microfluidic lab-on-a-chip devices are finally here and being used in the laboratory. Employing tiny structures to transport and control microscopic quantities of fluid, lab-on-a-chip devices are now commercially available from a number of companies for analysing biomolecules such as DNA, conducting protein crystallisation and performing simple chemical reactions. They may soon even be blasted into space to search for signs of extraterrestrial life.
But, in all likelihood, this is only the beginning. For a vast array of researchers around the world are hard at work trying to develop the next generation of lab-on-a-chip devices. These could progress beyond the laboratory and out into the everyday world, forming the basis for everything from novel medical diagnostics to tiny fuel cells. Such devices will need to be robust, easy to manufacture and a great deal more complex.

Stephen Quake, a professor of bioengineering at Stanford University, California, US, and one of the pioneers of commercial microfluidic chips, has likened the current situation to the early years of computing, before the development of the integrated circuit transformed it into a practical technology. At the moment, scientists are able to produce the kind of microscopic structures needed for microfluidics, but they need to find methods for joining these structures together into increasingly complex and integrated designs.
When this happens, microfluidics may begin to transform our lives to the same extent as computing. ’There’s the potential here for applications across almost every subject area,’ says Harp Minhas, editor of the RSC journal Lab on a Chip.
The proposed benefits of microfluidic chips are manifold. They offer the possibility of working with small samples and utilising tiny amounts of chemicals. They should produce faster and more consistent results than larger systems. They should also be easier to use, because all the necessary steps, such as sample preparation, mixing and separation, would be automatically conducted on the chip.
In order to realise these benefits, microfluidics researchers have already had to overcome a number of challenges. First off, they had to find ways of transporting fluids within the tiny channels (10-100
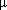
m wide) etched into chips.
The most obvious way to do this is simply to apply air pressure to one end of the channel using an external pump. But the problem is that, at these small scales, fluids tend to stick to the sides of the channels. This means that increasingly high pressures are needed to force the fluids through the tiny channels and these pressures can damage the chips.
An alternative method is to utilise a process known as electroosmotic flow, which commonly occurs when separating compounds by capillary electrophoresis (CE). When an electric current is applied along the length of a fluid-filled channel, it can cause negative ions to be generated on the channel wall, which in turn causes a layer of positive ions from the fluid to build up along the wall. These positive ions are then attracted towards the negative electrode, causing the fluid to travel in that direction. Electroosmotic flow thus offers a fairly straight-forward way of transporting fluids within tiny channels.
It’s therefore not surprising that miniature CE systems were one of the first microfluidic devices to be developed commercially, with analytical technology companies such as Agilent Technologies and Caliper LifeSciences now selling CE microchips. These comprise a simple system of microscopic channels (usually made up of a central separation channel and numerous side channels for adding the separation buffers and samples) and reservoirs etched into a glass or silicon chip. They are mainly designed for analysing biomolecules such as DNA and proteins and are already widely used in pharmaceutical research.
Chipping away
The very first glass and silicon microfluidic chips were developed by Andreas Manz and colleagues in the early 1990s, who pioneered the concept of micro total analysis systems (
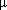
TAS). They used techniques borrowed from the microelectronics industry, such as photolithography and etching, which are precise but relatively expensive.
For more complex devices, however, glass and silicon are not suitable materials. Devices that require moving parts, such as valves, cannot easily be fabricated in such rigid materials. This has forced microfluidics researchers to turn to rubber-like polymers, in particular a silicone elastomer known as polydimethylsiloxane (PDMS).
This polymer has become the chip material of choice for microfluidics researchers. Ironically, this is not primarily because of its flexibility, but because complicated designs can be fabricated from it quickly, easily and repeatedly. For instance, one method known as hot embossing involves pressing a hot metal frame into the partially-molten polymer. In this way, a large number of identical microfluidic chips can be fabricated using a single frame.
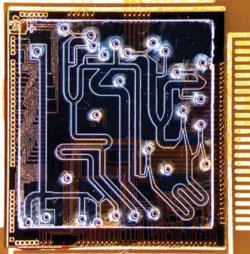
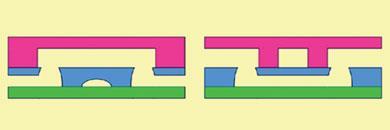
Another fabrication method for PDMS, known as ’multilayer soft lithography’, was developed by George Whitesides at Harvard University, Massachusetts, US, in the late 1990s to take advantage of the polymer’s flexibility. Quake refined the method to generate thin layers of PDMS with defined shapes - by spinning the polymer onto a mould - and then sticking the layers on top of each other. Quake found that if he fabricated two channels, placed one above the other and then applied air pressure to the top channel, it would bend downwards to close off the bottom channel, acting like a valve.
By fabricating a large number of microscopic channels and valves on a single chip, Quake showed that he could produce highly complicated designs, in which fluid and samples can be directed along specific pathways and confined in specific compartments by opening and closing the valves. Furthermore, placing three valves close together and operating them in sequence produced an on-chip pump to move the fluid and samples around.
In 2002, he developed a chip with over 2000 separate microvalves and used it to test individual bacteria for the expression of a specific enzyme. Then, in 2005, he developed a chip-based microreactor and showed that it could conduct the five sequential chemical processes required to synthesise a radioactive probe used in a medical imaging technology known as positron emission tomography.
’One of the main challenges in developing practical microfluidic devices includes finding a microfluidic platform which has a good balance between functionality, complexity and ease of fabrication,’ explains Jessica Melin, a member of Quake’s research team and director of the Stanford Microfluidics Foundry, which custom-manufactures microfluidic chips for universities and research institutes. ’You want a platform which allows versatile and highly integrated microfluidic functionality while not being too difficult to fabricate. A platform such as multilayer soft lithography with integrated monolithic valves strikes a good balance.’
The first commercial product based on this microfluidic platform was developed by a US company called Fluidigm, which Quake co-founded in 1999. This is a chip called TOPAZ, which conducts protein crystallisation and was first launched in 2003. The main advantages of TOPAZ are that it can work with small samples of protein and can simultaneously explore numerous different routes to crystallisation, in order to identify the most efficient one.
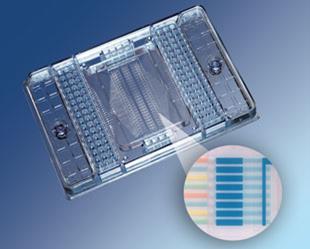

Similar PDMS microvalves and pumps are central components of a microfluidic device for detecting signs of extraterrestrial life, which has been developed by a team of researchers led by Richard Mathies, a professor of chemistry at the University of California, Berkeley.
’We have been working, in total, for nearly 10 years on the development of microfluidic technologies to detect very low levels of organic molecules that we think might be biomarkers on other planets,’ explains Mathies. ’Recently, we’ve developed some highly sensitive detection methods, which can detect amino acids down to parts per trillion.’
Mathies’ microfluidic device works by first heating extraterrestrial soil samples to vapourise any organic molecules that may be present. These molecules then condense onto a cold finger coated with a fluorescent dye that binds solely to amino acids. Finally, these amino acids are separated by capillary electrophoresis (CE) in a buffer containing cyclodextrin compounds, which separates the amino acids based on their chirality. Life on Earth tends to prefer left-handed amino acids, and researchers think that a similar asymmetry would be a good indicator of biological activity on other worlds.
Chips in space
The device will soon get a chance to search for amino acids on Mars, as it has been invited to form part of the scientific payload of the European Space Agency’s ExoMars mission, which is due to launch in 2013. It is also showing its worth in more terrestrial applications. Mathies is currently using a version of the chip to investigate the wide range of organic molecules present in alcoholic drinks, in order to identify those that cause hangovers.
He has also adapted the microfluidic technology to create a chip that can conduct all the steps - thermal cycling, sample purification and CE - required in the current standard method for sequencing DNA. This chip is currently being commercially developed by the US company Microchip Biotechnologies, which was set up in 2003 to commercialise leading-edge microfluidic technologies.

Despite these successes, there are still numerous challenges that need to be overcome before microfluidic chips can match the ubiquity of computer chips. ’The miniaturisation community still doesn’t fully understand fluidics at the micro- and nanoscale,’ says Minhas. ’Not all the relevant laws apply in the way we would expect them to when you scale down.’
For instance, a continuing problem is how to mix two different liquids at the microscale. One potential solution was developed by Whitesides and his colleagues, who managed to mix two fluids as they travelled along a microfluidic channel by etching a herringbone pattern of ridges on the channel’s floor. These rotate the flow in different directions, thereby mixing the fluids together.
An alternative method was recently developed by two chemical engineers from Texas A&M University, Arjun Sudarsan and Victor Ugaz, who found that, at high enough flow rates, two fluids travelling alongside each other would swap positions as they travelled round a 180? bend in a channel. By repeatedly splitting and recombining the fluid flow through four smaller channels, Sudarsan and Ugaz could achieve 90 per cent mixing between the two fluids.
Other researchers are investigating the transport of individual liquid droplets, which can be moved around on open surfaces rather than trapped in channels. Thomas McCarthy of the University of Massachusetts, Amherst, US, has shown that a droplet can be moved around by creating hydrophobic water-hating ’paths’ surrounded by even more hydrophobic ’curbs’ on a surface. The only force needed to move the droplet is gravity.
Another challenge is to develop standardised microfluidic components, which can be linked together into a whole range of complex designs. ’Ideally we would like to achieve a design methodology similar to that of the microelectronics industry, where component libraries can be used to design an integrated circuit,’ says Melin. ’Similarly, microfluidic designers could potentially use microfluidic components to design large scale integrated devices without needing to know the microfluidic details of each component.’

Melin thinks that Quake’s system of PDMS valves and pumps could form the basis for such a system. Recent work by Axel Scherer and colleagues from the California Institute of Technology, US, (where Quake conducted much of his research before moving to Stanford in 2004) has probably made this more likely.
Using multilayer soft lithography, Scherer and his team have created PDMS structures that they call ’vias’. These allow individual channels to move between different layers on a chip, so that a channel can cross over or under another one without interacting with it. By extending microfluidics into three dimensions, Scherer and his colleagues have moved closer to the goal of creating microfluidic chips with large numbers of components and highly complex designs.
But as well as solving these technical problems, microfluidic researchers need to start finding some new ’killer applications’ for their devices. ’To achieve greater acceptance for this incredible technology we need to be able to solve real problems, rather than just doing research that’s interesting,’ says Minhas. ’We need to demonstrate its application to solving real-world, hopefully high-profile, problems that no other technology can address.’
To find and solve these problems, however, microfluidic researchers need to talk to those who are affected by them. ’If you’re developing a point-of-care test for influenza, for example, then researchers should ideally talk to the people who will actually use this technology before developing it, or involve them in the process,’ says Minhas. Jan Eijkel, a microfluidics researcher from the University of Twente in the Netherlands, agrees: ’You need the input of biologists in order to tell you what they would like to do.’
It took computers just 30 years to progress from a niche technology to being a central part of our lives. With increasingly complex designs and a few killer applications, chances are that microfluidic chips will make the same journey in an even shorter period of time.
Jon Evans is a science writer based in Chichester, UK
Synthesis on a chip
Analytical devices may currently dominate the microfluidics field, but it is the development of microreactors that perhaps offers greater potential for synthetic chemists.
Microreactors are microfluidic devices used for running chemical reactions. They offer many of the same advantages as other microfluidic devices, such as enhanced speeds and low chemical requirements, but they also have another major advantage: ease of scale-up.
Synthetic strategies developed in the lab often need to be completely redesigned to meet the demands of synthetic strategies on a larger, commercial scale, explains Graham Sandford at Durham University, UK. But if chemists had microreactors that could be used fairly cheaply in the lab, scale-up would simply involve employing a greater number of microreactors running in parallel.
Sandford and emeritus professor Dick Chambers are currently leading a team that is developing microreactors for conducting direct fluorination reactions. Most recently, the Durham group has developed a microreactor consisting of up to 30 channels, and used it for reacting fluorine gas with a range of different substrates.
Conventional fluorination reactions use expensive reagents, and while fluorine gas is cheaper, it is tricky stuff to use at large scales. But in Sandford’s microreactor, the fluorine gas and liquid substrates react together very efficiently in the microscopic channels. The technique could be used for other gases too, he says: ’What we’ve developed can be a general purpose tool for anyone looking to do scale-out of gas-liquid processes’.
Other microreactors are already starting to reach the market. For instance, Thales Nanotechnology, a Hungarian microfluidics manufacturer, has launched a microreactor, known as the H-Cube, for conducting hydrogenation reactions. This was put to use in a recent synthesis of the natural product (?)-oxomaritidine in seven synthetic steps by Steven Ley’s group at the University of Cambridge, UK (see Chemistry World, October 2006, p11).






No comments yet