US researchers use click chemistry to measure cell proliferation
Researchers in the US have developed a new way to detect DNA synthesis in living cells by using click chemistry - the concept of reacting together two ’high energy’ molecules that ’click’ together efficiently under mild conditions. Experts believe the technique could lead to new ways of identifying cancer cells.
To see how fast cells are proliferating, biologists measure the rate of DNA synthesis by feeding them analogues of the natural building blocks of the molecule. These synthetic compounds become incorporated into the growing DNA chain and can be detected after the cells are killed.
The two most widely used analogues are [3H]thymidine, whose radioactivity can be measured in the fixed cells, or 5-bromo-2’-deoxyuridine (BrdU), which can be detected by specific antibodies raised against BrdU.
However, both have drawbacks. Tritiated thymidine produces only a weak signal and the technique is time consuming. The disadvantage of the BrdU method, on the other hand, is that the bulky antibody may not diffuse through large tissue samples to reach the cells of interest.
Adrian Salic of Harvard Medical School in Boston, Massachusetts, and his colleague Timothy Mitchison have overcome these hurdles by using click chemistry.
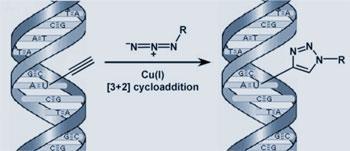
’We made a very simple chemical analogue of thymidine which contains a terminal alkyne in the form of an ethynyl group,’ Salic told Chemistry World. The triple bond of the ethynyl can be ’clicked’ with a fluorescent azide, a compound containing three double-bonded nitrogens in a row. This is a classic ’3+2’ cycloaddition - a rapid, efficient click-reaction catalysed by copper.
Cells fed the alkyne-containing analogue incorporate it into their DNA. The team then add the fluorescent azide, which quickly diffuses into cells and reacts with the ethynl group. ’You obtain a strong signal that is detectable under standard fluorescence microscopy at low magnification,’ said Salic.
The technique works in cultured cells and in whole animals. ’The method is foolproof,’ Salic added.
’The authors have used elegant, cutting edge chemistry to develop a novel approach to label replicating DNA in cells,’ John Diffley, deputy head of Cancer Research UK’s London Research Institute told Chemistry World. ’This work may lead to new diagnostic approaches for the early identification of cancer cells.’
Simon Hadlington
References
Proc. Nat. Acad. Sci., 2008, DOI: 10.1073/pnas.0712168105





No comments yet