First detailed structure of a key enzyme involved in cancer should help scientists develop new tumour-targeting drugs
Researchers in the US have produced the first detailed structure of a key protein involved in regulating cancer, an eagerly anticipated advance that should speed anti-cancer drug development.
Since its discovery in 1985, telomerase has attracted great interest for its roles in cancer and ageing. Now Emmanuel Skordalakes and colleagues at the Wistar Institute in Philadelphia have unveiled the fine detail of the enzyme’s catalytic site. They say their work will prove invaluable for drug developers. ’I feel this is going to open the door to the design of a host of molecules that hopefully will be useful cancer treatments,’ says Skordalakes.
Telomerase is responsible for adding protective DNA ’caps’ called telomeres to human chromosomes. In healthy adult cells, telomerase is deactivated as the body ages. Without the action of the enzyme, the telomeres get shortened every time the cell divides, eventually to the point that the cell can no longer replicate - a normal part of aging.
But in most tumour tissues, telomerase gets reactivated, making each cell effectively immortal because the telomere is continually repaired as the cells multiply out of control. As telomerase is active in most human cancers, but not in healthy tissue, it has been a key target for drug designers. However, the enzyme’s size and complexity have made its structure difficult to pin down.
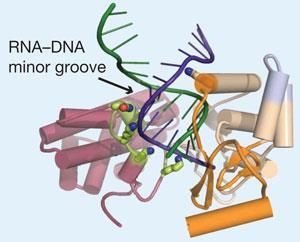
The team crystallised enzymes from red flour beetles, and used sophisticated X-ray diffraction techniques to build an electron density map of the catalytic protein, known as TERT, which they could then interpret using computer software. The DNA regions that code for functional aspects of the enzyme are well conserved across different species and, according to Skordalakes, the human version of the protein should look very similar.
’Now we can actually see how telomerase assembles together to carry out telomere synthesis,’ says Skordalakes. For instance, the team have already identified a number of amino acids that they think make direct contact with the substrate during the elongation process. One particular lysine has a side chain which may help to position the incoming DNA within the active site.
Comparisons of the model with other similar proteins support a common origin with HIV reverse transcriptases - enzymes that make DNA in HIV. ’It’s a really exciting and significant piece of work,’ says Stephen Neidle, head of the Cancer Research UK Biomolecular Structure Group at the University of London, UK. ’What they’ve shown is that the active site, as everybody thought, resembles the HIV reverse transcriptases, but there are differences and the differences are important to future discovery of TERT inhibitors.’
The anti-HIV drug azidothymidine (AZT) is already known to be active against TERT, but its effect is weak compared to its effect on HIV reverse transcriptases. ’Now we can tell why,’ says Skordalakes. ’With the three dimensional structure, we can sit down and look at existing inhibitors and see why they’re not working so well, so we can chemically modify them.’
Hayley Birch
Enjoy this story? Spread the word using the ’tools’ menu on the left.
References
A J Gillis, A P Schuller and E Skordalakes, Nature, 2008, DOI: 10.1038/nature07283






No comments yet