Using a molecular recognition approach to protein functionalisation allows amino acid side-chains to be modified selectively
Chemically modifying proteins is fundamental to biochemical research, but it is far from easy to get the result you want. Chemists in the US have now developed a powerful strategy for selectively modifying the side-chains of proteins, which they hope will enable the creation of new tools to investigate protein interactions involved in human diseases.
Modifying the side-chains of the amino acids that make up proteins is straightforward enough, but as a particular type of side-chain may appear many times in a given protein, modifying just one of them is a tough challenge. Brian Popp and Zachary Ball at Rice University, Houston, have been using the ideas of chemical reactivity and reaction design to try and solve this problem.
In their elegant solution, they decided to dispense with the standard approach of designing a highly selective reagent. Instead, they used a reagent that is hardly selective at all, but one that only works in the presence of a rhodium catalyst.
The clever bit is to place the catalyst exactly where it’s needed by attaching it to a coiled peptide that binds to the right bit of the target protein. This brings the catalyst and the side-chain into close proximity, allowing them to react as soon as the reagent - a diazo derivative of styrene ?- is added.
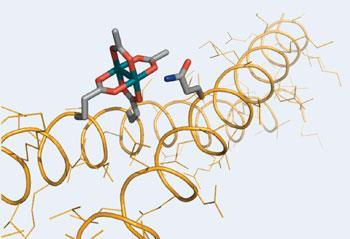
Ball thinks that this approach is ’a big step forward’. He says that its main benefit is that the high reactivity of the diazo compound allows it to react with the side-chains of over half the naturally occurring amino acids, ’a broader range than any established method,’ he says. The method is also highly specific, as any catalytic units that start to react where they are not wanted are quickly destroyed by reaction with water, which is the solvent for the reaction.
Ball says that as well as looking to establish the robustness of their new method, future work may involve investigating how it could be used to tag, image, or modify the structure or function of natural proteins. He adds, ’we believe this work will create powerful tools to investigate transient protein interactions, such as those along signalling pathways that lead to human disease.’
Carlos Barbas, Kellogg Professor of Molecular Biology at the Scripps Research Institute, La Jolla, US, is enthusiastic about the work, and says ’it’s a beautiful example of how peptide structure can be used to modify side-chain reactivity to allow for specific peptide labelling.’
David Barden
Link to journal article
Proximity-driven metallopeptide catalysis: Remarkable side-chain scope enables modification of the Fos bZip domainBrian V. Popp and Zachary T. Ball,?Chem. Sci., 2011, 2, 690DOI:10.1039/c0sc00564a





No comments yet