Computer program scores full marks in Cambridge contest
A team of researchers has hit the jackpot by correctly predicting the crystal structures of four organic molecules in a competition organised by the University of Cambridge.
The crystalline structure of a material defines many of its properties, including solubility, hardness and colour. Accurate prediction could be a boon for the pharmaceutical industry and help researchers design new materials with unique properties.
Cambridge’s ’crystal structure prediction challenge’, which has been running since 1999, requires teams to predict the crystal structures of four compounds from only the molecular structure and conditions of crystallisation. This year, seven of the 15 teams that took part were able to correctly predict at least some of the crystal structures using the latest computational techniques; and one team got all four right using a new computer program called GRACE.
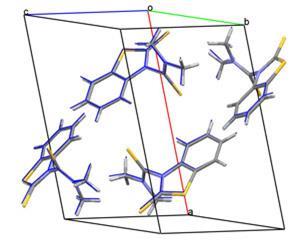
The winning team included Marcus Neumann from Avant-garde Materials Simulation (AMS) in France, along with Frank Leusen and John Kendrick from the Institute of Pharmaceutical Innovation (IPI) at the University of Bradford, UK. Neumann told Chemistry World: ’We can predict small molecules well, but it will take time to extend this to more complicated pharmaceutical molecules. We believe this could develop into a valuable complementary technology.’
The four compounds were chosen to have diverse properties, with different flexibilities, shapes and functional groups. They ranged in size from 10 to 25 atoms - smaller than most pharmaceutical molecules but big enough to be challenging.
Graeme Day at the Cambridge Crystallographic Data Centre, who organised the challenge, said, ’Last time (in 2004) we had a similar number of groups but only one correct prediction. It has been much more successful this year, partly due to some innovative new approaches and increases in computer processing power. But there is still a long way to go before we will see desktop-based solutions.’
In the future, the work could help the drug industry to solve the problem of polymorphs, where the same drug compound can form different crystal structures with different properties - and different effects on patients. Different polymorphs can also be patented separately, so finding them all is important for drug makers.
The next challenge is scheduled for 2010.
Lewis Brindley





No comments yet