Helicos sequences a viral genome with technology that could help to make personalised medicine a reality
The prospect of treatments that are tailored to fit an individual’s genetic makeup is a step closer thanks to technology unveiled by US scientists. A team at Helicos Biosciences in Massachusetts have pioneered a fast, cheap way to read long stretches of DNA - and demonstrated that it works by sequencing the genome of a virus.
Mapping out the sequence of the 3 billion base pairs that make up the human genome is currently an expensive and time-consuming process. Conventional ’Sanger sequencing’ has dominated the field since its conception 30 years ago and works by replicating DNA strands in the presence of fluorescently-dyed nucleotides. As the chains grow, dye molecules can enter in place of normal nucleotides and prevent the chain from growing further.
By using four different fluorescent dyes that correspond to the four different nucleotide bases and measuring the lengths at which the chains stopped growing, the sequence can be deduced. But in order to get accurate information, large amounts of genetic material are needed. This is provided by enzymes that copy the DNA thousands of times.
Now the Helicos team have demonstrated the potential of their method - ’true single molecule synthesis’ (tSMS) - by accurately reading the 6400 base pair long genome of the M13 virus.
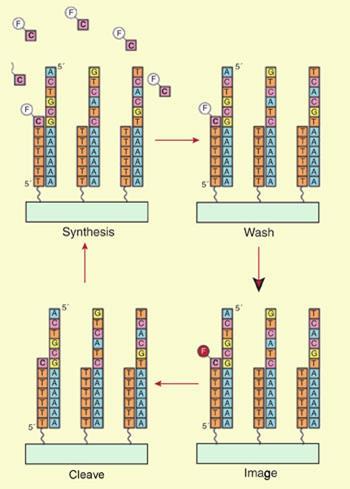
The tSMS technique does not require the DNA to be amplified - making the process cheaper and potentially more accurate as replication can allow errors to creep in. It works by attaching tails of a long repeating nucleotide to fragments of DNA, anchoring them onto a surface around 100 nanometres apart - a distance that allows the strands to be imaged separately.
’We can then read off the sequence by adding a single nucleotide at a time,’ explains Tim Harris, senior director of research at Helicos Biosciences. ’We image a single dye label on a single nucleotide that is added to a single strand - then we add another one.’
One by one, the sequence can be built up on the DNA strand, providing direct access to sequences of more than 25 bases for each strand. By using a specially designed imaging camera, thousands of individual strands can be imaged at once, gradually allowing a picture of the genetic code to be built up.
’This is an elegant demonstration showing that single molecule sequencing has arrived,’ says Steven Block, an expert on DNA sequencing at Stanford University, California.
’There is still some way to go before this can be used to map human DNA, but it is really a matter of scale,’ Block adds. ’If the technique can be used to measure 6000 bases there is no reason why it couldn’t potentially measure 3 billion in the future.’
Lewis Brindley
References
et alScience, 2008, DOI: 10.1126/science.1150427






No comments yet