Measuring how small molecules interact with structures such as telomeres could throw up cancer drug targets
A new single molecule technique can provide greater insight into the interactions between G-quadruplexes and small-molecule ligands. These insights could give novel perspectives for cancer drug testing and design say the researchers.
G-quadruplexes are formed from guanine (G)-rich portions of nucleic acids - DNA and RNA - and are often found at the end of telomeres, the pieces of DNA that cap the end of chromosomes that have been implicated in cancer cells’ immortality. This means that G-quadruplexes are potential anti-cancer drug targets and a great deal of research is being put into investigating how ligands interact with these structures and interfere with telomere replication.
The international team from universities in the US, UK and Japan have worked together to create a technique which fixes a G-quadruplex between two beads, and then measures how much force it takes pulls those beads apart. Small molecules that stabilise the quadruplex may be interesting drug targets worth following up.
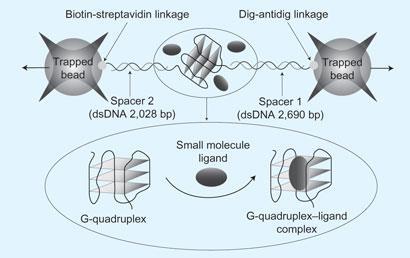
Previous work has provided a lot of structural information and insights into the folding and unfolding kinetics of quadruplexes but mechanical information on how much ligands can stabilise the structure has been lacking. The new work provides data about all three of these variables.
’Mechanical forces are important to consider because there can be competition between polymerase-mediated transcription/replication processes and the ability of the polymerase to "read through" G-quadruplexes that are stabilised by ligand binding,’ says Nathan Luedtke from the University of Zurich, Switzerland. But, in the body, G-quadruplexes aren’t unfolded by pulling at the ends of DNA, but selectively unfolded by specific enzymes.
’This is an interesting paper because most of the previous studies have focused on ensemble measurements in thermodynamic equilibrium. Living systems are, however, not in thermodynamic equilibrium,’ Luedtke adds.
Laura Howes
References
D Koirala et al, Nat. Chem., 2011, DOI: 10.1038/nchem.1126






No comments yet