Researchers pinpoint enzyme and mechanism responsible for producing polycyclic compounds with anticancer and antibiotic activity
US researchers have worked out how some microbes produce polyketides - a class of polycyclic compounds that have antibiotic and anti-cancer properties but are difficult to manufacture. The finding could open the way to manipulating the system by genetic engineering to produce new drugs.
Microbes are thought to produce polyketides by first making a long-chain linear polyketone then bending and folding it to produce the polyaromatic molecule. But precisely how they accomplished this feat of origami was a mystery.
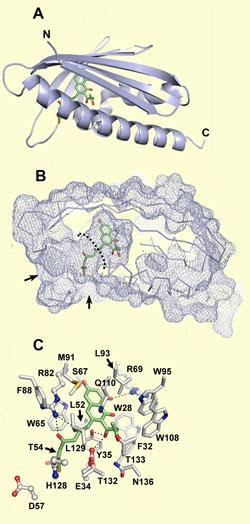
Now, Sheryl Tsai and colleagues at the University of California at Irvine have worked out the crystal structure of a bacterial enzymecalled aromatase/cyclase, which was known to be involved in the process but whose exact role was unclear. The researchers showed that the protein contains a C-shaped active site perfectly configured for housing a long-chain linear polyketone and the various intermediate structures that are produced as parts of the chain are successively aromatized. Theoretical analysis of the site showed that it has the relevant functional groups to hold the molecule in place and perform the appropriate chemistry.
The researchers hope that by subtly changing aspects of the active site they will be able to control the size and shape of the polyketide, resulting in the formation of new drugs.
’Almost every polyketide has rings in its chemical structure, and if we can control ring formation, we can produce more polyketide drugs,’ says Tsai. ’Until now, polyketide ring formation was a mystery that hampered our efforts to produce new drugs.’
’These long-chain polyketones are extremely reactive and would otherwise react spontaneously to form a whole range of different products,’ says Russell Cox, a polyketide biosynthesis expert at the University of Bristol in the UK. ’The enzyme appears to be able to physically restrain the molecule and configure it very selectively without making any mistakes.’
Simon Hadlington
References
B Ames et al., Proc. Nat. Acad. Sci., 2008, DOI: 10.1073/pnas.0709223105





No comments yet