Large organic molecule mimics biological protein folding
French researchers have synthesised a large organic molecule that folds up like a small protein, though its backbone is entirely non-biological. The achievement is a step along the path to producing truly synthetic enzymes in the laboratory.
The scaffolding of biological proteins, made up of just twenty naturally occurring amino acid types, folds into complex structures - like helices and sheets - that determine the protein’s overall function. Over the last decade, organic chemists have tried to copy these structures artificially by synthesising other families of folding molecules, collectively called foldamers, which don’t use biological amino acids in their backbones.
So far, foldamers have not approached the size or complexity of a biological protein, said Ivan Huc of the European Institute for Chemistry and Biology, Bordeaux, France, who led the research team. Small molecules have been made which mimic sub-elements of large proteins - like a single helix - but few have shown interactions between two sub-elements, corresponding to a protein’s tertiary structure.
Now Huc’s team has synthesised and obtained the crystal structure of a large non-natural molecule, made of over a thousand atoms, which folds up into two helical units connected by a short linker. The helices are forced close together, and influence each others’ twisting direction.
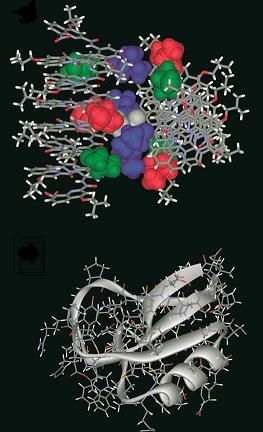
This structure might not seem much of an advance over a single helix. But, as Huc pointed out, two helices give the new foldamer a complex surface shape, with an inside and an outside. If decorated with carefully positioned catalytic groups, it could begin to behave rather like a natural enzyme.
’This is beautiful work, typical of the elegant and pioneering efforts from the Huc laboratory,’ said Sam Gellman, who studies foldamers at the University of Wisconsin-Madison, US. But Gellman also suggested that the helices’ forced interaction was different from the types of tertiary interactions seen in biological proteins.
The researchers prepared their foldamer by attaching branched molecules together, rather than by building up a sequence of small monomer units, as protein machinery does. ’It is a masterpiece of synthesis and control over the supramolecular structure of a complex molecular system,’ said Bert Meijer, who works in macromolecular and organic chemistry at Eindhoven University in The Netherlands.
Huc expects his team, and others, to construct more complex foldamers in the future. ’Making a truly synthetic enzyme is still a huge challenge,’ he said.
Richard Van Noorden
Putting some backbone into bacterial killers
Molecules need a bit of backbone in order to punch through bacterial membranes, say US chemists.
A polyamide that folds back on itself, biting its tail, to make a compact three dimensional structure has been stumbled across by researchers in the UK
References
N Delsuc et al, Angew. Chem. Int. Ed., 2006, DOI: 10.1002/anie.200603390





No comments yet