Next Generation Sequencing has been with us for little more than a decade, but has already revolutionised biological science. According to Jonas Korlach, chief scientific officer of US-based sequencing company Pacific Biosciences (PacBio) we are now entering a golden age of sequencing technology and the advanced understanding of genetics it brings; ‘Now that we have those tools to really move forward with that understanding, we can use that knowledge to improve human health.’

The reduction in cost and speed of sequencing has fuelled an expansion of its use. Some analysts predict the global market will grow from $5.9 billion (£4.3 billion) in 2020 to $23 billion by 2025. When Illumina’s vice president, Mark Ross, joined the company in 2007, ‘the possibility that you could generate a whole human genome sequence in just a couple of months was a huge revelation’. He remembers sceptically asking colleagues if they thought the technology could be improved. ‘It’s proven to be a very naive question,’ he jokes.
He puts Ilumina’s success down to its ‘beautifully elegant method,’ and the R&D investment the company makes: in 2020, Illumina invested $682 million, which is about 21% of its revenue. This has so far reduced the cost of generating a human genome to $600, as well as improving accuracy and the length of individual DNA fragments that can be read.
But there are new sequencing methods taking on Ilumina’s market dominance. In 2013, ThermoFisher Scientific acquired Ion Torrent for $16 billion. The non-optical method is more stable then other systems says Andy Felton, vice president of product management at Ion Torrent. It is semiconductor-based and detects hydrogen ions. ‘You can think of it as a really tiny pH meter,’ says Felton. A polymerase incorporates natural nucleotides onto DNA fragments in millions of tiny wells and a change in voltage detected for each addition. ‘The advantage of the technology is we can do many more reads in the same timeframe,’ says Felton.
Long-read sequencing
As with Illumina, Ion Torrent is a short read technique – with a maximum read length of about 400 bases. ‘We can generally have a crack at sequencing almost anything,’ says Felton, but it does pose problems for highly repetitive and duplicated genome regions because it is difficult to find unique sections to compare to reference samples. Some medically relevant genes fall into this category, including the Human Leukocyte Antigen (HLA) system, which codes for parts of immune function. ‘People typically are using long read sequencing more for those,’ says Michael Quail, who is involved in evaluating sequencing instruments at the Wellcome Sanger Institute, near Cambridge, UK. ‘Like a jigsaw, in small bits is very difficult to put together, but a jigsaw in large bits is easier.’

A new generation of sequencers are able to read longer DNA segments – upwards of 10,000 bases. Until recently these technologies tended to be error-prone. But, says Korlach, ‘about two years ago, we broke the paradigm of having two worlds, one with short, accurate reads, and one having long noisy reads, by taking the best of the two and creating what we call HiFi reads.’ PacBio’s long read Single Molecule, Real-Time (SMRT) Sequencing technology was co-invented by Korlach and Stephen Turner, both at Cornell University in New York, US. The company moved to California and went public in 2010.
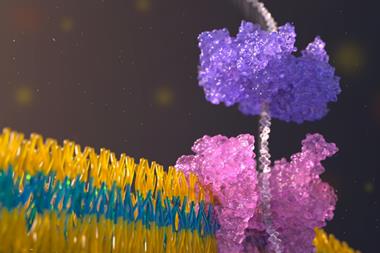
The optical technology immobilises single DNA molecules in tiny wells called zero-mode waveguides. ‘We’re watching each DNA polymerase separately and you can watch it for as long as it goes, because you don’t have to keep all the [amplified copies] synchronised,’ says Korlach. It’s made possible by synthesising nucleotides with the fluorescent label attached to the terminal phosphate rather than the base. ‘Now [when] the polymerase cleaves the phosphodiester bond [it] separates the label from the incorporated nucleotide,’ explains Korlach, ‘after each incorporation cycle, the polymerase has no knowledge that there was a fluorescent tag involved and that ensures an efficient process.’
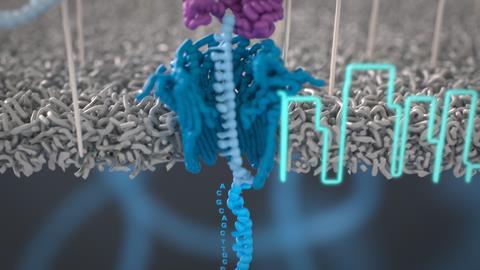
The other major long-read technology comes from Oxford Nanopore, originally developed by Hagan Bayley at the University of Oxford, UK, and has a record read of around 4 million bases. It also follows single DNA molecules, trapped in nanopores embedded in an electro-resistant membrane. As DNA moves through the pore the current is disrupted to produce a characteristic ‘squiggle’ which can be read in real time to determine the DNA or RNA sequences of multiple molecules. ‘[This] means that sequencing occurs much quicker and individual molecules can be sequenced within several minutes,’ says Quail. As well as an ability to create long reads, in 2014 Oxford Nanopore released its first portable ‘pocket-sized’ sequencing device.
Although the human genome was first sequenced in 2001, hundreds of gaps had persisted in areas of ‘structural variation’, including repeated and inverted motifs that may be responsible for disease. They are mostly invisible to Illumina short read sequencing, but in April the Telomere-to-Telomere (T2T) consortium completed the full sequence of human chromosome 8.1 ‘PacBio technology enabled this feat,’ and says Korlach ‘we have just begun to uncover all of the variation that exists.’
Prize-winning science
‘We didn’t aim to tackle sequencing, we got funding from the [Biotechnology and Biological Sciences Research Council] to do a very basic experiment, ‘ explains chemist David Klenerman. Along with University of Cambridge, UK, colleague Shankar Balasubramanian, they were trying to watch real-time DNA synthesis using fluorescently labelled nucleotides. ‘We realised that the experiment we were doing could basically be changed very slightly to enable us to do very rapid sequencing.’ Ultimately the Next Generation Sequencing (NGS) technology they developed has provided a million-fold improvement in sequencing speed and cost and a revolution in the biological sciences.

Balasubramanian and Klenerman are keen to point out they did not do this alone, but with a large and interdisciplinary team of talented people. ‘We did a lot of proof-of-concept work in the chemistry department,’ says Balasubramanian. The method, known as sequencing by synthesis (SBS), starts by fragmenting DNA into many small single stranded pieces. These are immobilised in a flow cell and copied using fluorescently tagged colour-coded nucleotides with a reversible terminator that blocks incorporation of the next base until the colour is detected, after which the terminator is cleaved. ‘The power of [the method] is that you can array molecules and parallelise the whole process,’ says Balasubramanian.
Balasubramanian and Klenerman set up Solexa in 1998. In 2006, they released their first commercial genome analyser, which could sequence billions of bases in one experimental run. One change from the original design was to add an amplification step to improve accuracy by multiplying the DNA fragments. The bridge amplification method they chose came from Pascal Mayer (honoured alongside Klenerman and Blasubramanian in the 2022 Breakthrough prize) and Laurent Farinelli and was pivotal to Solexa’s success.
Solexa was acquired by Illumina in 2007 and Balasubramanian says the company must take credit for hugely improving the system since then, now sequencing trillions of bases per experiment. Although there were other high-throughput DNA sequencing methods, Illumina quickly dominated the market. In 2015, the company claimed over 90% of the world’s sequencing data was generated using Illumina instruments.
Klenerman says they knew their work would have an impact, but they didn’t predict the scale; ‘it’s quite a surprise how widely it is [now] used’. The surprise for Balasubramanian is the range of uses; ‘there are now literally hundreds of applications, [all] ways of detecting counting and mapping different features in biological systems using a sequencing readout as a tag or a surrogate. For me, this was unimaginable at the outset.’
One advance that Balasubramanian thinks will be crucial in the future is the ability to sequence epigenetic markers on DNA, such as methylation. ‘I think, going beyond the four genetic letters will provide a more comprehensive picture of what’s going on in biological systems,’ he says, In 2012 he launched Cambridge Epigenetics, which is developing technology for directly sequencing methylation and other base modifications.
‘We’ve gone through a quantum leap,’ says Balasubramanian, ‘a small lab can now do what the whole world couldn’t do 20 years ago. But we have to keep going because there’s more information that we can get from DNA.’
Cancer diagnostics
For most NGS companies, the future lies in developing new applications. One of the discoveries that has paved the way is the presence of small DNA fragments in blood. First used for non-invasive pre-natal testing of foetal DNA, companies have now moved on to cancer diagnostics. Ross, who is part of Illumina’s medical genomics research group, says the firm now sees itself as an oncology diagnostics developer. ‘We don’t focus just on the sequencer, we have a strong focus on the end-to-end process.’
Illumina’s TruSight Oncology 500 kit provides clinicians with information on mutations in about 500 genes found in solid tumours. The aim is to make sequencing-based diagnostics easy enough to be carried out by non-specialists. Ion Torrent is also developing this approach. ‘We envisage that we can get next generation sequencing to be simple enough, both from the technical wet side operation, and the bioinformatics, for a majority of labs who are involved in pathology workflows to be able to run it,’ says Felton. The latest system requires only 10 minutes of hands-on time to get from sample to report.
Ultimately for NGS to become routine and available to all cancer patients, ‘[we need] more clinical trials in which genome sequencing is compared with [the] standard of care, to show that there is a clinical, as well as economic benefit,’ says Ross. ‘Certain healthcare systems are starting to embrace human whole genome sequencing, ’he adds, pointing to NHS England’s Genomic Medicine Service, which aims to sequence NHS patients with cancers and rare diseases.
Infectious disease monitoring – the Covid-19 challenge
Cancer diagnostics is not the only application for NGS of course. ‘If there are any silver linings around Covid-19, I think it has shown what sequencing can do,’ says Korlach. ‘Traditionally, illumina has had less of a focus on infectious disease.’ But, says Ross, ‘the pandemic has changed that situation dramatically’.
Illumina developed kits for Covid-19 surveillance, creating a reference genome against which other viral genomes could be compared. ‘That’s critical, because it allows us to have a surveillance program in which we can understand different variants of the virus, how they are spreading in communities,’ says Ross. In 2020, Illumina donated $1.4 million of sequencing systems and consumables to ten African countries to support surveillance efforts across the continent.
Ion Torrent has also moved into infectious disease surveillance over the last two years. ‘We’ve increased the sensitivity pretty significantly…we know that we’re going into an era of more vaccinated and asymptomatic patients so we’re going to have low[er] viral loads.’ Its equipment can sequence a full viral genome from as little as 20 viral copies. Oxford Nanopore developed the LamPORE assay, a cheap and highly scalable Covid-19 diagnostic test, which detects the virus in 90 minutes and is being used by the UK government for rapid screening of healthcare workers. Unlike most other assays, LamPORE sequences RNA and amplifies three highly conserved genes in Sars-CoV-2 positive samples.
The way forward
Today’s sequencing capabilities are a work in progress according to Korlach, ‘there’s still a long way to go for us to get to a point where sequencing becomes so ubiquitous, comprehensive and accurate, that we would say, we are done!’ Quail jokes that we haven’t yet invented Star Trek’s ‘tricorder,’ which could analyse a species in a matter of seconds.

Despite the huge cost reductions (in 2007 it cost $300,000 to sequence a human genome, compared to $600 today) Felton says cost is still ‘the biggest barrier we see to wider adoption’. In 2017, Illumina’s chief executive, Francis D’Souza, announced his ambition to reach the $100 human genome mark. This would then ‘democratise the use of genomic sequencing in health care,’ says Ross. The possible applications at that point are immense. For example, Quail imagines we might use sequencing as a tool for rapid bedside diagnostics of meningitis, or to instantly identify antimicrobial resistant bacterial strains. Others envisage sequencing all babies at birth, although Ross acknowledges ‘there are obviously ethical considerations that need to be discussed and addressed in that area.’
‘It’s highly likely that you’re going to have a blood-based cancer test in the next five to 10 years that is going to detect the early presence of cancer, and that’s going to be routinely used to monitor your health,’ concludes Felton, and this is likely to be possible for other diseases too. It may be that in the future genome sequencing will be a part of regular medical check ups, where mutations and epigenetic changes can be tracked through life. But for these advances we may need to wait a little longer, until our ability to interpret the human genome catches up with the data that sequencing technologies can already provide.
References
1 G A Logsdon et al, Nature, 593, 101, 2021 (DOI: 10.1038/s41586-021-03420-7)














No comments yet