An ancient enzyme has been reconfigured to catalyse a non-natural reaction at a speed comparable to that of an average modern enzyme towards its natural substrate.1 Precambrian β-lactamase, as the name suggests, catalyses β-lactam hydrolysis and can degrade β-lactam antibiotics, but thanks to a new computational tool researchers have engineered it to have high activity in an anthropogenic reaction called a Kemp eliminase. ‘This is significant both from a general design perspective, and also that this significant improvement in activity was obtained through pure computational screening, which opens significant doors to future enzyme design studies on challenging systems,’ says Lynn Kamerlin of Uppsala University in Sweden, who led the work.
More and more chemists are turning to enzymes in a bid to make their reactions more sustainable and precise. Modern enzymes have evolved to be highly specific so are difficult to repurpose. Ancient enzymes, however, were more promiscuous, so starting with older enzymes gives researchers more scope to deploy them for non-natural reactions.
Kemp elimination involves proton abstraction from carbon centres. The reaction acts a benchmark for artificial enzyme design, since no natural catalysts have evolved for the process. Kamerlin and her colleagues have previously reconfigured Precambrian β-lactamases for a Kemp elimination reaction using a rational design strategy.2 And though that work demonstrated how extinct enzymes could act as scaffolds for enzyme engineering, the resulting enzymes weren’t better than existing Kemp eliminases.
Now, using software called FuncLib alongside empirical valence bond calculations the team has designed an enzyme with a turnover number of ∼102s−1. The enzyme’s activity is only one order of magnitude lower than the most efficient Kemp eliminase, which itself took multiple rounds of directed evolution to create.
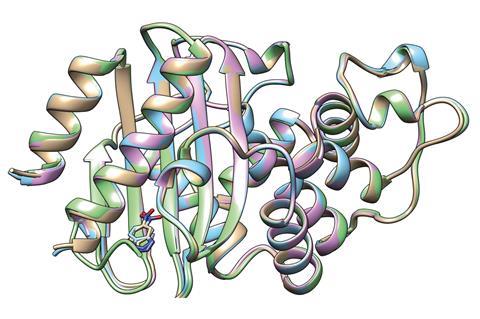
Directed evolution is a protein design method that exploits Darwinian evolution. While the technique has revolutionised protein engineering, it doesn’t work so well for non-natural enzymes where the active site has often been designed independently from the rest of the structure. ‘Directed evolution is costly and requires many rounds of evolution to see substantial gains in activity,’ says Kamerlin. ‘Computational approaches have been rapidly advancing to be able to bridge this gap, but have so far been quite far still from what can be achieved through directed evolution.’
‘We believe FuncLib is a game-changer that is going to substantially accelerate the enzyme design process,’ says Kamerlin. FuncLib makes changes to an amino acid sequence and performs evolutionary analysis and stability calculations to predict the best enzyme variants. As it focuses on stability, the researchers added empirical valence bond calculations to test for activity. Kamerlin explains that empirical valence bond calculations are much cheaper than making the enzymes experimentally, so act as a way to narrow down the selection. They could therefore select for both stability and activity, increasing the chances of finding a highly active variant. Although FuncLib has been proven for natural enzyme design this work shows that FuncLib also works for non-natural reactions.
Valerie Welborn, an expert on quantum chemical simulations of complex condensed-phase systems at Virginia Tech in the US is impressed that FuncLib works on synthetic enzymes. ‘It is easy to see that stable multipoint mutants computed with FuncLib can serve as a better starting point for directed evolution. This does not only save tremendous resources by speeding up the directed evolution process, it also provides a new route for cases where countless rounds of directed evolution previously failed to provide satisfactory results.’
References
1 V A Risso et al, Chem. Sci., 2020, DOI: 10.1039/d0sc01935f (This article is open access.)
2 V Risso et al, Nat. Commun., 2017, 8, 16113 (DOI: 10.1038/ncomms16113)










No comments yet