Microfluidics researchers are aiming to bring new diagnostic devices into mainstream medicine. Mark Peplow reports

Handheld devices that carry out medical tests in minutes are a transformational technology. Rather than shipping millilitre samples of blood or urine to centralised laboratories for processing and analysis, point-of-care diagnostics offer rapid answers from microlitre samples with minimal preparation. These microfluidic lab-on-a-chip technologies could be particularly useful in a range of situations: emergencies, helping patients monitor their own health, or in places with little clinical infrastructure.
Over the past two decades, microfluidics researchers have developed thousands of miniaturised biomedical assays, and the global point-of-care diagnostics market has boomed to more than $17 billion (£10.5 billion), according to a recent estimate.1
Yet lab-on-a-chip devices are still a long way from realising their full potential. Earlier this year, an analysis2 by David Beebe, a biologist at the University of Wisconsin, Madison in the US, and colleagues showed that some 85% of studies in this field are still published in journals focused on engineering, physics or chemistry, rather than biomedicine – indicating that they were about development, rather than implementation of the devices.
‘We have journals and journals of papers on this kind of thing, but clinical diagnostics has not actually changed very much at all over the last several decades,’ says Shana Kelley at the University of Toronto, Canada, who works on microfluidic diagnostics.
There are many reasons for this mismatch, including the difficulties of winning regulatory approval for diagnostic devices and the huge sums of money required to take any product from bench to clinic. But in many cases, lab-on-a-chip devices have simply not offered enough advantages to shift away from traditional testing systems. ‘To be successful, it has to win on multiple counts – not just cheaper, not just faster, not just a better test – it has to be a combination of all of these to give you an advantage in the clinic,’ says Beebe.
A growing band of researchers is now finding new ways to address these hurdles. Rather than focusing on one aspect of a miniaturised assay, they are taking a more holistic approach: producing finished, user-friendly devices and spinning out companies to turn them into a commercial reality.3 ‘Researchers are really shifting how they do things,’ says Samuel Sia, a biomedical engineer at Columbia University in New York, US. ‘They’re recognising the challenges.’
A drop in the ocean
The most ubiquitous point-of-care devices in today’s market tend to rely on lateral flow tests, where analytes wick through a paper strip, picking up labeling reagents that ultimately produce a visible band in a readout area. The technology is used in home pregnancy tests and to detect flu infections and HIV. In contrast, the glucometers used by people with diabetes rely on electrodes to measure the enzymatic breakdown of glucose, offering a precise blood sugar reading.
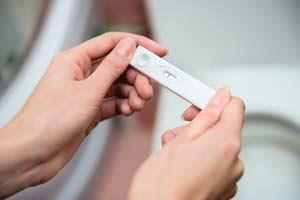
These are enormously successful products, but they illustrate the limitations of current devices, and the difficulties that more complex assays must overcome. Lateral flow tests are cheap, but they typically only detect one biomarker at once, are not particularly sensitive, and offer qualitative results that are open to misinterpretation. In some cases, lateral flow tests may not be cost-effective because they result in too many unnecessary treatments, or even misdiagnoses. ‘False positives and negatives are very expensive,’ says Aaron Wheeler, an analytical chemist at the University of Toronto (interview: Algae-on-a-chip).
Glucometers offer a quantitative readout, but they detect an analyte that is found at a much higher concentration in blood than most other disease biomarkers, making it much easier to get an accurate reading. The sheer number of people with diabetes, and the frequency with which they need to test their blood, also creates a huge market, which helped to attract investment to develop the devices.
More sophisticated devices that can tackle more than one biomarker at once are also seeing greater use. The i-STAT, for example, is a handheld blood analyser that can measure ions such as sodium and potassium, blood gasses, haemoglobin and more. Its manufacturer, Abbott, says that about 50,000 of the devices are now in use, carrying out 100 million tests per year. ‘It’s become the standard frontline system in emergency rooms,’ says Wheeler.
Nevertheless, after winning FDA approval in 1992, the i-STAT ‘took a long, long time to penetrate the market’, says Beebe. ‘And compared to the potential market for point-of-care diagnostics – which is everyone, including patients, doctors and pharmacists – it’s a drop in the ocean,’ says Sia.
Pump up the volume
To expand their reach, point-of-care devices need to be completely automated, requiring virtually no sample preparation. That means picking out very low concentrations of particular biomarkers from a soup of similar molecules in untreated blood or urine.
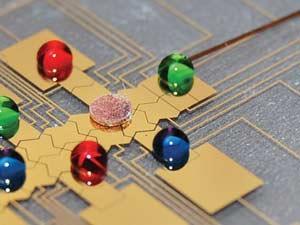

Traditional laboratory assays may use dozens of steps to purify their samples and then amplify the signals of key biomarkers. Take Elisa (enzyme-linked immunosorbent assay), one of the most common laboratory tests for biomolecules such as peptides, proteins or hormones that provoke an immune response – broadly classed as antigens. During the test, target molecules are tagged with complementary antibodies that are connected to an enzyme. This enzyme causes a colour change in a colourimetric reagent, which flags the presence of the target biomarker. The test can also be used to detect antibodies themselves.
An automated point-of-care device that simply recreated the conventional laboratory Elisa would be bulky, complex and expensive. So Sia has developed an alternative, which he calls the mChip.4
Instead of using enzymes, his assays tag biomarkers with antibodies connected to gold nanoparticles and then treats them with a solution of silver ions. Wherever the antibodies have bound to their target, the silver can be detected by a system of LEDs and photodetectors that together cost a few pounds.
The tests are carried out on microfluidic cassettes made from transparent polystyrene, which cost about six pence each and can be produced from a mould at a rate of one every 40 seconds. Reagents move around the chip contained within discrete droplets to avoid cross-contamination; samples can even be pulled through the device by the vacuum formed by drawing a syringe from its barrel.
Sia’s team has used the mChip to detect antibodies against HIV and Treponema pallidum, the bacterium that causes syphilis. Both are treatable in pregnant mothers, to reduce the likelihood of them passing the diseases to their offspring. ‘My goal is to get this product to the developing world,’ he says.
The researchers have tested their device on hundreds of patients in Rwanda, finding it to be as sensitive as lab tests and about ten times faster.5 Crucially, the mChip can catch weakly positive samples that would be missed by the lateral-flow HIV test. They have even incorporated satellite and mobile phone transceivers so that data can be transmitted directly to a centralised medical facility. ‘It’s a nice case study of trying to do it right,’ says Beebe. ‘Until you start using samples from a patient, you don’t really know if it works.’

Sia points out that much of the basic technology used by the mChip is more than a decade old. He founded a spin-out called Claros in 2004 to commercialise the system, which was bought by multinational pharmaceutical and diagnostics company Opko Health in 2011. It has since won approval in Europe for a prostate cancer test using the system.
But Sia has continued to refine his design. All too often, he says, researchers in the field focus too much on developing novel assays, rather than uniting existing technologies into a viable device. ‘It’s an integration challenge,’ says Sia. ‘All the pieces are there – but how do you put them together?’
Cheap as chips
Sia’s choice of material – polystyrene, chosen for easy manufacturing – highlights another stumbling block for commercialisation. Historically, lab-on-a-chip diagnostic devices have tended to be made from glass, silicon or polydimethylsiloxane (PDMS). ‘PDMS has arguably been the most popular microfluidic device material – but it’s not cheap, it’s not ideal for biological work and it’s difficult to scale up manufacture,’ says Beebe.
So researchers are increasingly turning to materials that are more amenable to high-volume, low-cost production. Sia went with plastic; but Wheeler and many others are now working with paper microfluidic devices that are cheap, disposable and easy to pattern by inkjet printing.
PDMS has been the most popular material – but it’s not cheap
David Beebe
Complex, multistep assays can be trickier to implement on paper, partly because of the possibility that residual reagents will contaminate each other. So Wheeler corrals his samples and reagents within discrete droplets that may have volumes of just nanolitres – a technique called digital microfluidics.6 Electrostatic fields generated by an array of inkjet-printed electrodes move droplets around the chip, merging or splitting them as needed so that they can go through multiple procedures without needing pumps or other moving parts. Wheeler estimates that each device takes about one minute to print and costs less than three pence. And since the channels are not carved into the chip, changing the electrostatic fields generated by the electrode array can create a new route map for the droplets. ‘It’s easily reconfigurable,’ Wheeler says.
His team has set up several assays on these chips, including a proof of principle test for a rubella antibody, IgG, which appears after someone has already had a rubella infection, or has been vaccinated against it.7 It requires 30 separate steps that take just 10 minutes, and is as reproducible and sensitive as conventional tests – something that a simpler lateral flow test would be unable to do, he says. They’re now working on combining this with a test for another rubella antibody, IgM, which indicates an ongoing infection and is difficult to distinguish from IgG. In unpublished results, his team has been able to measure both antibodies as accurately as conventional lab tests, and his spin-out company, Kapplex, aims to commercialise the work.
Let it slide
Chemists and engineers have sometimes assumed that clinicians would be prepared to learn new techniques – fluid handling, for example – in order to use microfluidic devices. ‘But to move them out of centralised labs and closer to patients, these assays have to be completely automated,’ says Kelley.
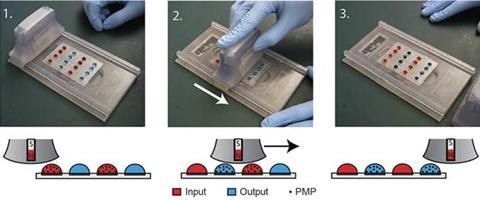
Beebe’s lab has focused on simplifying sample processing. His Slide (sliding lid for immobilised droplet extractions) technology relies on microscopic paramagnetic beads to extract nucleic acids and other biomolecules from sample droplets. Paramagnetic beads have long been used in this way in the lab, but it involves a lot of solution transfer and wash steps that have been too complex to reliably incorporate into point-of-care devices.
Slide contains a sliding handle fitted with magnets that lifts the beads out of the droplets and moves across the device like an old credit card imprinter. The handle transfers the beads – and their cargo of analytes – to a second set of droplets containing analytical reagents, where magnets in the base pull the beads away from the lid.
The researchers have refined the design so that there is virtually no carryover of liquid from one bead to another, making the tests as accurate and reliable as conventional assays. Earlier this year, Beebe reported that beads coated with a specific antibody could reliably separate green fluorescent protein from red fluorescent protein, as well as DNA from viral RNA.8
Beebe is developing the device through his spin-out company Salus Discovery, in partnership with other companies. ‘It will be commercialised very soon – it’s the culmination of all our sample preparation work,’ he says.
Close to the clinic
Beebe reckons that microfluidic researchers have too often focused on over-engineering their devices, perhaps creating a diagnostic that is too complicated to use or difficult to manufacture. The solution, he says, is to work in close collaboration with those who are likely to be the end users – biologists and clinicians – and focus as much effort on making the devices simple to use as adding functionality.

Five years ago, Beebe moved his lab to a hospital, and he credits this with making a huge difference to his research. ‘I’m reminded every day that the reason we’re doing this is because of the patients,’ he says. It’s also easy to talk to clinicians on an almost daily basis, quickly finding out whether particular data could improve treatment.
It’s a lesson that Kelley has also taken to heart in Toronto. She is tackling one of the toughest problems in point-of-care diagnostics – identifying trace nucleic acids – and relies on collaboration with clinicians to identify useful tests and to source patient blood samples for her work.
During her PhD at the California Institute of Technology in the US, she helped to develop a way to use electrochemistry to find mutations in a DNA sequence, and ever since then she has been improving that system, a neutraliser displacement assay that she calls AuRA.9
The devices contain tiny electrodes that are nanostructured to give them a huge surface area, and coated with single strands of negatively charged DNA. These DNA probes have just the right sequence to bind specific analytes such as DNA, RNA or proteins.
20 minutes can make a huge difference
Shana Kelley
The negatively charged probe DNA is initially bound to a peptide nucleic acid bearing cationic amino acids, which neutralises its charge. But when an analyte binds to the DNA probe it displaces the neutraliser, causing a measureable change in charge at the electrode. That electrical signal would be too weak to reliably detect trace biomarkers, so Kelley has added some cunning redox chemistry to amplify the signal.
Freed of its neutraliser, the probe DNA’s negative charge attracts a positive ruthenium complex from solution, initiating an electrochemical cycle that ultimately creates a much larger electrical change at the electrode. This enables very sensitive and specific detection of analytes within minutes, even from unpurified samples.
Kelley says that they can achieve the same levels of sensitivity and specificity as traditional DNA tests without having to use enzymes to amplify the signal, which makes the system a lot easier to manufacture and package. In 2010, Kelley founded a company, Xagenic, to bring the technology to market. Their first product, the X1, is aimed at diagnosing infectious diseases. ‘That’s where 20 minutes can make a huge difference,’ she says. Xagenic plans a clinical trial next year and – pending regulatory approval – it hopes to launch the device in 2016.
Kelley says that more and more researchers working on point-of-care diagnostics are recognising the need for simplicity, manufacturability and marketability. ‘There’s a lot more automation being put into diagnostic systems now – chemists sometimes think that’s just basic engineering, but it’s really important,’ she says. ‘It’s what lab on a chip is all about – going from sample to answer without having a human running around a lab, but instead having a fluid running around a chip.’
It’s a change in approach that will be instrumental in realising the full potential of point-of-care microfluidics, enabling doctors to routinely perform molecular diagnostics in their offices. ‘It’s going to happen,’ says Kelley. ‘Five years from now, that’s going to be standard practice.’
Mark Peplow is a science journalist based in Cambridge, UK
References
1 The point of care diagnostics market forecast 2014–2024, VisionGain, 2014
2 E K Sackmann, A L Fulton and D J Beebe, Nature, 2014, 507, 181 (DOI: 10.1038/nature13118)
3 C D Chin, V Linder and S K Sia, Lab Chip, 2012, 12, 2118 (DOI: 10.1039/c2lc21204h)
4 C D Chin et al, Nat. Med., 2011, 17, 1015 (DOI: 10.1038/nm.2408)
5 C D Chin et al, Clin. Chem., 2013, 59, 629 (DOI: 10.1373/clinchem.2012.199596)
6 M Abdelgawad and A R Wheeler, Adv. Mater., 2009, 21, 920 (DOI: 10.1002/adma.200802244)
7 R Fobel et al, Adv. Mater., 2014, 26, 2838 (DOI: 10.1002/adma.201305168)
8 B P Casavant et al, Anal. Chem., 2014, 86, 6355 (DOI: 10.1021/ac500574t)
9 J Das et al, Nat. Chem., 2012, 4, 642 (DOI: 10.1038/nchem.1367)







No comments yet