While Pfizer and Moderna request emergency approvals for their coronavirus vaccines, researchers in Hong Kong and the US have discovered a new weapon that could fight the virus. Combining de novo design and directed evolution, they created a protein that binds to Sars-CoV-2 spike protein, blocking its infection pathway. The protein neutralises the virus in vitro and shows promising results in hamsters, protecting them from coronavirus infection after a small intranasal dose.
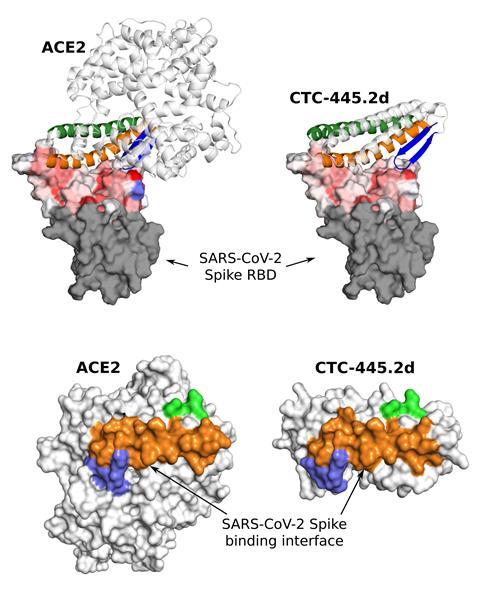
Infection by Sars-CoV-2, the coronavirus that causes Covid-19, takes place after the virus’ spike glycoprotein binds to ACE2 receptors on the surface of our cells. ‘We have now created a decoy ACE2 protein that binds to the coronavirus spike really strongly,’ explains Daniel Adriano Silva, from Neoleukin Therapeutics, who led this study. The firm’s proprietary technology allows them to first analyse an existing biomolecule, identify key structures and functions, and then create computational simulations for proteins with similar behaviour. Neoleukin can then design a protein to bind that target. ‘Our team has created a protein with binding domains like the ones in ACE2,’ adds Silva. ‘We trick the virus into binding to them and neutralise it.’
The process involved a gargantuan effort, both computational and experimental. ‘We started with a set of 100,000 protein designs, of which we selected 35,000 candidates that offered good binding,’ comments Silva. After a careful selection process, the team decided to synthesise 198 proteins, and then tested their binding affinity to Sars-CoV-2. The best candidate only shares around 30% of its sequence with human ACE2 and is five times smaller. However, the important part remains the same – the binding domain is identical, and that is enough to trick the virus.
‘This is key to avoid a negative immune response,’ says Chris Bahl, an expert in protein design at the Institute for Protein Innovation in Boston, US. ‘Using the smallest possible amount of the human protein minimises the risk of triggering an autoimmune disease, and their design strategy hinders the ability of the virus to develop resistance to the treatment.’
Once researchers had identified the best candidate, they improved its affinity and specificity using directed evolution. ‘This is a technique that mimics the process of natural selection by intentionally introducing mutations,’ explains Jim Montclare, an expert in protein engineering at New York University, US. In this case, the team used a polymerase enzyme that introduces random mutations during transcription. Then, if any variants bind better to the target, they can be identified and characterised, explains Montclare. In this case, directed evolution yielded a protein with nanomolar affinity to Sars-CoV-2 – an improvement on the original candidate.
Both Bahl and Montclare are impressed by these results. ‘They developed a protein that recognises Sars-CoV-2 spike protein with high affinity and specificity from scratch,’ says Montclare. ‘The speed and fidelity of the process is the true breakthrough; the team created a decoy protein that works extremely quickly,’ says Bahl – the whole process took less than 10 weeks.
The first experiments, both in vitro and in vivo are promising. Silva and his team administered a single intranasal dose of the protein to hamsters, and then gave them a lethal dose of Sars-CoV-2. ‘Normally, they would be really sick after seven days, but with a prophylactic dose of our protein all of them survived,’ he explains.
De novo proteins are still new to the therapeutics scene. ‘They have just been around for a couple of years, and Neoleukin is among the first companies looking into clinical trials,’ says Bahl. The technology is promising, though – the proteins could be easily produced at scale using cell cultures. Bahl believes the current vaccines and monoclonal antibodies will suffice to tackle Covid-19, ‘but this tool could be really useful to fight future pandemics’.
References
TW Linsky et al, Science, 2020, DOI: 10.1126/science.abe0075
















No comments yet