Nanopore DNA sequencers are on the verge of becoming a reality, as controlling DNA strand movement through the pore is finally cracked
Rapid, cheap DNA sequencing has just edged a step closer, thanks to a new technique to control the motion of single DNA strands as they pass through a protein nanopore ’reader’. The advance, which was the work of Reza Ghadiri and his team at the Scripps Research Institute in La Jolla, US, cracks the fundamental problem that has been holding back nanopore-based DNA sequencing - how to get the DNA to travel slowly enough through the pore to read off the sequence base by base.1
The idea behind such protein pore-based readers - such as those recently developed by Hagan Bayley at Oxford University, UK2 - sounds simple in principle. An electrical potential is used to draw single DNA strands from one side of the pore to the other. As each strand enters the pore, the current flowing through the pore changes measurably - and the exact current is characteristic of the DNA base occupying the pore at the time.
In theory, the DNA sequence can then be read off by recording the changing current as the DNA strand traverses the pore. However, in practice, the voltage needed to thread the stand into the pore then sucks the DNA through the pore far too quickly for each individual base to be detected.
Ghadiri’s solution is to attach a polymerase enzyme - the protein nature uses to turn single-stranded DNA into double-strands during cell replication - to the top of the DNA strand. As the enzyme works its way along the strand, adding bases one by one to form the second strand, it gradually draws it through the pore, base by base at a rate that would be slow enough for each base to be electrochemically identified.
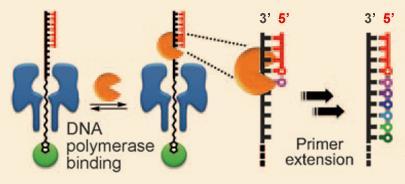
’You can think of the polymerase as a motor protein,’ says Ghadiri. ’It’s not just slowing the speed of DNA going through the pore that is important - you have to go through the pore in this stepwise fashion.’
’This remarkable demonstration that DNA can be ratcheted through a nanopore by an enzyme, without the cyclic addition of reagents, complements recent successes in base identification,’ says Bayley, who collaborates with Ghadiri on DNA readers but was not involved in this latest research. ’When the two are combined, a nanopore sequencing device will be complete.’
Ghadiri says he is already working on the next generation version of his system, which will combine his DNA polymerase approach with the kind of ’read-head’ nanopore developed by Bayley that can identify each base as it passes through the pore. ’It is another challenge putting all these things together, but at least there is no fundamental issue that it cannot be done.’
James Mitchell Crow
References
1. J Chu et al, Angew. Chem. Int. Ed., 2010, DOI: 10.1002/anie.201005460
2. J Clarke et al, Nature Nanotechnol., 2009, 4, 265 (DOI: 10.1038/nnano.2009.12)






No comments yet