Researchers have adapted mass spectrometry to image the distribution of molecules in tissues or membranes.
Updated 01 November 2006
Three research groups have made remarkable progress in using mass spectrometry for imaging biological tissue. Two have obtained very high resolution information on the distribution of molecules in isotopically-labelled samples, while the third has imaged unadulterated biological samples, albeit on a larger scale.
Steven Boxer’s group at Stanford University, US, used a commercially available NanoSIMS (secondary ion mass spectrometry) instrument to map the distribution of different lipids in an artificial membrane. In this technique, the surface to be studied is bombarded with a highly focused beam of caesium ions (Cs+), resulting in a flood of small secondary ions chipped off the target molecules.
’The NanoSIMS50 offers the combination of very high spatial resolution and very high sensitivity,’ said Boxer. While this technique can resolve down to around 50 nanometres, the secondary ions it produces are typically small fragments, and not very specific to their original biological molecules. In order to make the instrument distinguish between different kinds of membrane lipids, the group had to use an artificial membrane1 composed of two kinds of lipid labelled with the stable isotopes 15N and 13C.
The secondary-ion method has also been used by Claude Lechene of the Harvard Medical School in Cambridge, Massachusetts, and colleagues, to develop multi-isotope imaging mass spectrometry (MIMS).2 This allowed them to study a range of biological tissues that have been ’doped’ with different isotopes - for example, by feeding isotope-enriched amino acids to mice. His technique can achieve a lateral resolution of 33 nanometres, cope with samples just a few atoms thick, and distinguish between ions of very similar mass, such as 12C15N and 13C14N.
Lechene believes that this will allow scientists to track groups of molecules within a cell, or monitor specific cells within a group.
Both techniques give us a new window on biology because the spatial resolution of optical techniques is generally restricted to a few hundred nanometres, while atomic force microscopy probes down to the molecular level. Since these mass spectrometry techniques can resolve the organisation of biomolecules at length scales of 10 to 100 nanometres, they neatly fill this gap in our imaging capabilities, explained Jay Groves of the University of California, Berkeley. What’s more, they deliver both structural and chemical information about the sample.
No chemical alteration of any kind is needed in a different mass spectrometric imaging method developed by Graham Cooks’ group at Purdue University in West Lafayette, Indiana. Cooks’ approach relied on the desorption electrospray ionisation (Desi) technique, which he has been developing over the past few years. ’The novelty of this experiment,’ explained Cooks, ’is that it allowed, for the first time, ionisation of organic and biological compounds from samples which were freely accessible in the ambient environment.’
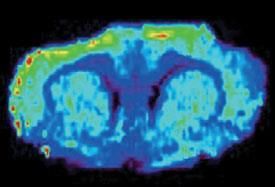
In earlier work, Cooks’ group traced one-dimensional profiles across biological samples, determining the changes of molecular composition along the way. Now they have expanded this approach into two dimensions, recording images of tissue sections. The resulting images can also show the relative concentrations of molecules in the tissue.3
While this method operates at a resolution of around a quarter of a millimetre, its high sensitivity for chemical differences without the need for labelling means it could be used to analyse tissue during surgery to establish tumor margins, said Cooks.
Carol Robinson from the University of Cambridge, UK, is impressed by the studies. ’This is a really exciting development. By analysing tissues and lipid bilayers without destruction, these researchers introduce a whole new dimension - that of spatial resolution,’ she said. Michael Gross
6 July 2005: Mass spec ensures crime doesn’t pay
Heroin-contaminated banknotes can be identified quickly and cleanly using tandem mass spectrometry, report chemists at the University of Bristol, UK.
References
et al, Science 313 , 1948 et al, J. Biol., et al, Angew. Chem. Int. Ed





No comments yet