Experiment identifies proteins from their mechanical fingerprints
Researchers have invented nano-sized calipers to identify individual protein molecules, which could make it easier to find trace proteins in biological samples. ‘We want to do for proteins what next-generation sequencing did for DNA,’ says Wesley Wong from Harvard University, US, who developed the technique with his colleagues.
Identifying individual proteins can be a tricky business. Proteins can’t be quickly replicated like DNA, so if you only have a tiny sample to work with there is no good way to get more of it. A technique called mass spectrometry is the gold standard for protein identification, but it only really works when there is a large amount of protein to work with.
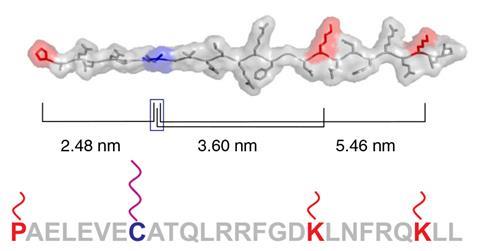
Researchers led by Wong and his Harvard colleague William Shih developed a set of tiny calipers to measure the nanoscale distances between different parts of a single protein. Those measurements act like a fingerprint to identify the molecule.
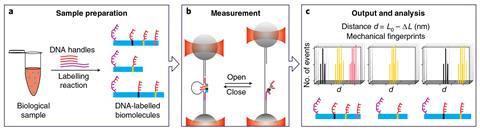
It works by attaching small tags made of single-stranded DNA to specific amino acids along the length of an unfolded, linear protein. The caliper jaws are made of short stretches of single-stranded DNA complementary to the protein tags, with a length of DNA stretched between them. When a protein is placed between the jaws, DNA pairing between the caliper and the protein tags creates a DNA loop.
As the researchers pull the jaws apart with beams of light – optical tweezers – or magnets, the protein-bound tags are released and the loop opens up, allowing the researchers to measure the distance between two tags. By making several measurements in that way and comparing them to a protein sequence database, the protein can be identified.
The technique could allow scientists to determine which proteins are being expressed in a single cell, which could be useful for diagnosing and profiling tumours. And if the team is able to extend the method to study proteins’ 3D shape, it could have applications in drug discovery, helping scientists understand how a protein changes when it binds to another molecule.
Chirlmin Joo, a biophysicist at Delft University of Technology in the Netherlands, says the technique shows a lot of promise, especially once the researchers work out how to use it on 3D proteins. ‘One challenge is how to do this on a large scale, to stretch thousands to millions of different proteins simultaneously, so it can analyse complex proteomes in a reasonable time scale,’ says Joo.
Wong says that his team is already working on scaling the approach to work on many molecules in parallel. ‘It will require some creativity, but we have a number of ideas,’ he says.
References
P Shrestha et al, Nat. Nanotechnol., 2021, DOI: 10.1038/s41565-021-00979-0









No comments yet