A nanomechanical mass spectrometer has accurately measured the mass of a huge 105MDa DNA-filled viral particle. The system can detect single molecules at a mass range missing from the suite of common mass spectrometric techniques.
The first mass spectrometer developed in 1919 measured the mas of atoms to detect differences in the number of neutrons in their nuclei. Over the past century, the technology has advanced to measure increasingly large molecules. Commercial instruments can now measure polymers and proteins as large as 1MDa, while mass spectrometers modified in research labs can work with molecules up to tens of MDa. Even more massive molecules are difficult to measure, however, because their sluggish ions resist deflection toward a detector by an electric or magnetic field.
A team led by Christophe Masselon and Sébastien Hentz, at the French Alternative Energies and Atomic Energy Commission, wanted to create a system to measure the missing mass range: MDa to GDa particles, the size of many viruses, large protein complexes and disease biomarkers. ‘We tried really to depart from the classical design of a mass spectrometer to find something that was really suited to deliver particles to a nanomechanical detector,’ Masselon says.
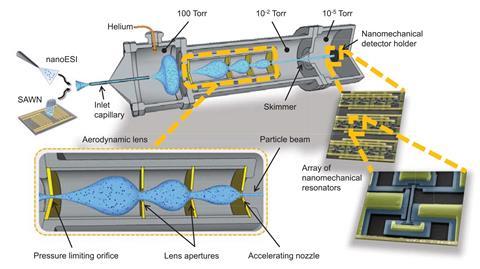
First, the researchers continuously ionised a solution of DNA-filled T5 viral particles using nano-electrospray ionisation. Once inside the instrument, the ionised droplets mix with helium gas. The mixture then flows through an aerodynamic lens, a low pressure chamber with a series of three openings that use the particles’ inertia to focus them into a tight beam.
As the mixture passes through each opening, the helium quickly diffuses into the chamber while the relatively sluggish larger particles diffuse more slowly. The particles’ paths become increasingly condensed as they pass through each opening, and the paths remain close together because the particles do not diffuse quickly.
Finally, the focused particle beam hits a detector tailored for this mass range, an array of 20 nanomechanical resonators etched into a silicon chip. Each resonator has a tiny beam suspended between two anchors. When a virus lands on a vibrating beam, the frequency of the beam’s oscillations change predictably based on the particle’s mass and location.
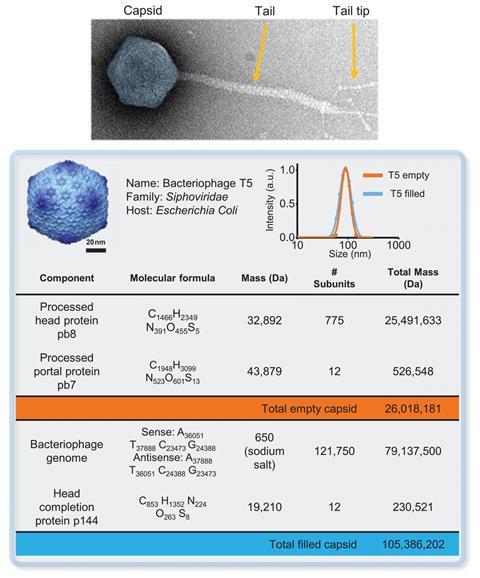
The researchers measured hundreds of DNA-filled viruses and found that the normalised distribution of measured masses centered on 108.4MDa. This value is slightly higher than the calculated molecular mass of 105.4MDa, perhaps because of salt introduced during ionisation, Masselon says.
The aerodynamic lens in this system is a nice way to introduce large species, particularly exotic ones like viruses, into a mass spectrometer, says Michael Roukes at the California Institute of Technology. He first developed nanomechanical mass spectrometry and worked with Masselon and Hentz on early instrument designs. Roukes sees the future of nanomechanical mass spectrometry as a tool to measure individual intact protein complexes from complex mixtures, identifying each component in a cell to detect low abundance disease biomarkers.
The design of this new system is also useful, Masselon says, because it can be easily modified to use other types of nanomechanical detectors, such as ones that detect particle shape, size and stiffness.
References
S Dominguez-Medina et al, Science, 2018, 362, 918 (DOI: 10.1126/science.aat6457)











No comments yet