Study builds scientists’ arsenal against drug-resistant superbugs
Scientists in the UK, Belgium and the Netherlands have gained a crucial understanding of the structure–activity relationship of new antibiotic, teixobactin. Since reports of its discovery in early 2015, researchers have shown it can kill a number of pathogens without them developing resistance to it.
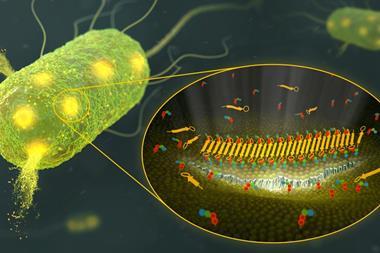
The University of Lincoln’s Ishwar Singh explains that there are several reasons for teixobactin’s potency: ‘It uses multiple modes of action to kill resistant bacteria, this makes it very attractive since, if it worked by only one mode, bacteria could modify more easily. It is much more challenging for bacteria to mutate on multiple levels.’ Teixobactin also targets lipids in the bacteria’s cell walls, which are considered to be less able to mutate and develop resistance.
Now, Singh and his team have synthesised and studied the three-dimensional molecular structure of seven teixobactin analogues using NMR and computer simulations, and discovered that the antimicrobial activity of teixobactin directly correlates with its molecular structure.
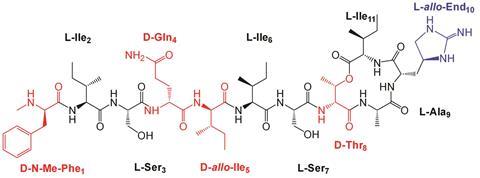
Teixobactin is composed of four D- and seven L-amino acids. By changing the D-amino acids to their L-counterparts, Singh’s team identified the role of the key D-amino acids, and their influence on both the molecular structure and activity of the antibiotic. They found that changing some of the residues from D to L resulted in a more ordered hairpin-like structure, which weakened its antimicrobial properties. D-residues introduced greater disorder into the system, which was found to be key to its antibiotic performance. The more disordered the structure is, the fewer intramolecular interactions there were between the molecule’s residues, freeing them up to bind to cell wall lipids, inhibiting cell wall synthesis and killing bacteria.
Since D-amino acids are more expensive than those with an L configuration, understanding which portions of the molecule are key for antibiotic activity will enable scientists to develop forms of the molecule that are both effective and financially viable. With an estimated 480,000 people contracting multi-drug resistant tuberculosis annually, this research is a step towards clinically marketable superbug resistant antibiotics.
James Nowick, an expert in the field of protein-inspired molecular design and synthesis at the University of California at Irvine, US, describes the work as a valuable contribution. ‘By understanding how different portions of the molecule contribute to its activity, we can design new molecules with enhanced beneficial features, and fewer detrimental features,’ he says.
References
This article is free to access until 16 March 2017
A Parmar et al, Chem. Commun., 2017, DOI: 10.1039/c6cc09490b








No comments yet