New method to hunt down protease-stabilised peptide inhibitors of large proteins
Researchers in the US have identified natural and unnatural peptides that bind Ebola virus glycoproteins using a new simple microarray. The positive peptide hits could form the basis of new Ebola therapies.
Regulators have approved more than 60 peptide-based drugs and hundreds more are in clinical trials and preclinical development. Peptides can be good therapeutics as they are generally selective and safe and well tolerated. Peptides, however, tend to be susceptible to digestion by the body’s protease enzymes. A way to get round this is to substitute in unnatural amino acids (D-amino acids rather than the usual L form), which are more resistant to protease. Nevertheless, hunting down therapeutic peptides made from D-amino acids is tricky and one of the most successful strategies for finding them only works on small and medium protein targets.
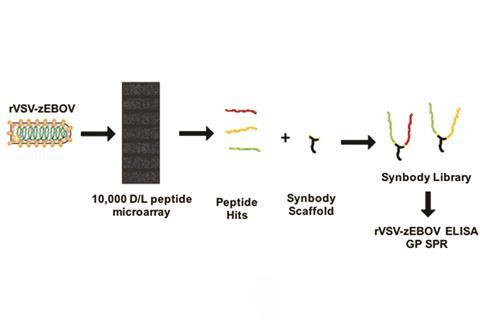
Now, Chris Diehnelt and his colleagues at Arizona State University have developed a platform to screen protease-stabilised peptides against large complex protein targets like viral glycoproteins. To use the platform against Ebola the team engineered a vesicular stomatitis virus pseudotyped with the glycoprotein from the Zaire strain of Ebola (rVSV–zEBOV). This concept maintained the glycoprotein’s tertiary structure but meant the researchers didn’t need the maximum biosafety facilities required for handling Ebola. So the team used their platform to screen rVSV–zEBOV against D and L peptides to find D-amino acid containing peptide binders for Ebola virus glycoproteins.
Diehnelt then constructed a library, which consisted of different duos of peptides hits from the microarray, joined by a scaffold. Screening of this library using ELISAs (enzyme-linked immunosorbent assay) for ones that bound rVSV–zEBOV was the final step in finding the potential therapeutic candidates.
‘Unlike other technologies which search large libraries, 108 to 1014 size, to find high affinity binders, we search small libraries and build affinity through bivalency [using two rather than a single peptide]. The discovery process is rapid and is quite robust,’ explains Dienhelt. He goes on to state that the peptides are mostly unstructured so are less sensitive to storage temperature – a clear benefit over the monoclonal antibodies used in Ebola drugs such as ZMapp.
Peptide expert Wuyuan Lu from the University of Maryland, US, believes this research could be a potentially important contribution to the peptide drug discovery field. However, he also notes that ‘due to its preliminary nature – in the absence of stability and bioactivity data – much remains to be done before its full potential can be realised.’
Diehnelt says his future plans involve optimising the activity of these ligands to develop broadly neutralising ligands that can inhibit the Ebola virus as well as other filovirus glycoproteins. ‘The ultimate goal is several high affinity, broadly reactive peptide ligands that can neutralise multiple filoviruses.’
References
This article is free to access until 22 February 2018
J A Rabinowitz et el, Chem. Commun., 2018, DOI: 10.1039/c7cc08242h













No comments yet