Chemists in Denmark have for the first time imaged chemical reactions on a DNA origami scaffold so that they can precisely attach single molecules
Chemists in Denmark have for the first time imaged chemical reactions on a DNA origami scaffold so that they can precisely attach single molecules. Their method, which involves atomic force microscopy, will help researchers to create self-assembling DNA nanodevices for applications ranging from biosensing to drug delivery.
Although DNA itself has limited potential in chemistry, optics and electronics, its structure can serve as a template for building materials with new functional properties. For complex structures, long DNA molecules can also be folded into different shapes, known as DNA origami. The idea is that different molecules can then be attached to the origami ’pegboard’ so composite structures become tailored, say, for conducting or optical circuits.
The trouble is that while larger nanoparticles or biomolecules destined for DNA can be imaged, smaller sub-molecular groups cannot. The standard imaging technique uses an atomic force

microscope (AFM), in which a vibrating probe detects atomic features through changes in resonant frequency, and its resolution is too poor to detect some parts of molecules involved in chemical reactions. In the past, this has prevented scientists from pinpointing where certain reactions take place, which in turn has prevented them attaching single molecules precisely.
Now, however, Kurt Gothelf and colleagues at Aarhus University have found a way around this problem. ’It is the first demonstration of chemical modification of a folded DNA nanostructure, and it holds promise for using DNA origami as a nanofactory for constructing advanced organic molecules,’ said Gothelf.
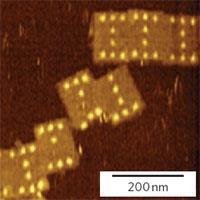
Using the new technique, the Aarhus team attach strands of oligonucleotides to the functional groups that are to be incorporated into the DNA origami pegboard. These ’staple strands’ will later bind to specific areas on the origami, allowing attachment with nanometre-scale precision. The team then tack the small molecule biotin on to the staple strands, and add the large protein streptavidin, which binds strongly to the biotin. Streptavidin can be imaged using AFM, and thus reveal the location of the groups on the DNA pegboard. As chemical reactions will cause the biotin-streptavidin complex to attach or be removed, the process can be easily observed using AFM as the complexes appear as bright protrusions in the images.
’This is neat stuff,’ said Hao Yan, an expert in DNA nanostructures at Arizona State University, US. ’It is a new way to control chemical reactions. Their demonstration that chemical reactions can be spatially localized and visualized on a DNA origami pegboard opens a lot of doors to implement potential molecular assembly lines.’
Gothelf told Chemistry World that his group are now looking to create chemical reactions between different points on the DNA pegboard, rather than at single sites.
Jon Cartwright
References
N V Voigt et al, Nature Nanotechnology, 2010, DOI: 10.1038/NNANO.2010.5




No comments yet