If you’re rich in Australia you’re more likely to drink plenty of coffee and eat citrus fruits, while if you’re poorer you’ll have less fibre in your diet and be more likely to be on painkillers. That’s a couple of the findings from one of the first studies that has attempted to link wealth to the kind of chemicals and dietary markers you can find in sewers.
Many wastewater studies have focused on chemicals as environmental pollutants. In this study, the researchers showed how differences in amounts of drugs and similar chemicals in wastewater can be linked to population demographics.
‘Our study is novel because we triangulated the per capita loads of chemicals in wastewater with the socio-demographics of the people from which the wastewater originated,’ explains lead author Phil Choi of the University of Queensland in Australia. ‘It’s an exploratory approach to investigate factors linked to different consumption behaviours. Our study is also the first peer-reviewed study to show that aspects of diet can be measured in wastewater.’
Choi and his colleagues analysed wastewater from six Australian states over seven consecutive days in 2016 looking for specific biomarkers. Then they used data from the Australian Bureau of Statistics to look for patterns.
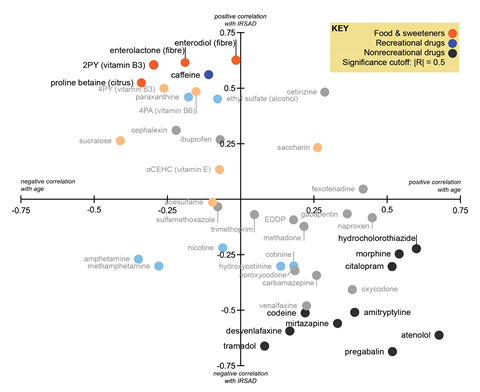
They found that coffee, alcohol and vitamin B markers were higher in areas of cities where more wealthy people lived. Most antidepressants and opioids, as well as the beta-blocker atenolol used to treat high blood pressure, anti-convulsant pregabalin and painkiller tramadol, appeared at higher levels in areas with low socioeconomic status. Consumption of citrus and dietary fibre was lower in areas where there were more people who did not finish high school or whose highest education was vocational training, compared with people in managerial positions or who attended university. However, socioeconomic status did not appear to be a factor in the levels of antibiotics or artificial sweeteners people consumed.
‘It is not easy to get this sort of fine sociodemographic resolution of data,’ Choi explains. ‘Using traditional, survey-based methods, one would need millions of people to objectively and consistently record their diet and sociodemographic information. Wastewater analysis provides an additional dataset, one which can potentially be continuously collected in near real-time to understand chemical consumptions/exposures of entire populations in a more objective and cost-effective manner.’ Choi believes such data could make health interventions more targeted and effective. ‘For example, our methods could be used to identify areas where diet could be improved, and subsequently to monitor the efficacy of targeted programs.’
This study takes the wastewater field beyond the illicit drug realm and moves it into new areas, comments Daniel Burgard, professor of chemistry at the University of Puget Sound. ‘This novel approach will hopefully introduce the power of the technique to folks doing epidemiological studies looking for new data sources.’ However, this approach has uncertainties, he points out. ‘It is unable to determine whether levels found in the wastewater are from high use by a few people, or low use by many. But this is also a benefit because it remains anonymous data. Survey-based data can provide information about individual consumption; wastewater can provide information about population level consumption.’
There are many challenges in interpreting data obtained this way, agrees Arjun Venkatesan, associate director for drinking water initiatives at Stony Brook University in the US. ‘The results can be biased if the sewage received flow from industrial discharges, runoffs from storm events and improper drug disposal in sewage. Also, the chemical can degrade or get transformed in sewer lines. [But] careful calculation and limitations of the data is often provided by the researchers.’ However, he thinks the approach is an extremely valuable tool for getting data that cannot be obtained through traditional surveys.












No comments yet