Understanding how chemistry links RNA triplets to the properties of amino acids
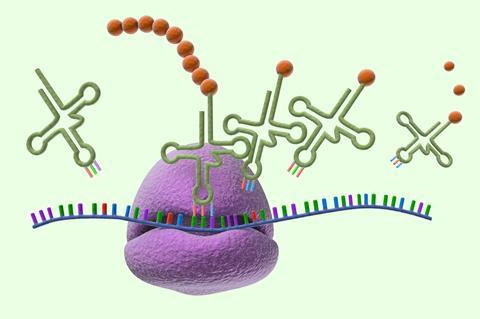
The genetic code has been dubbed ‘life’s greatest secret’.1 The code relates a sequence of DNA nucleotide bases in a gene to that of amino acids in the protein that gene encodes. Each of the 20 amino acids in proteins is represented by a different triplet of the four DNA nucleotides (denoted C, G, A, T) – or more properly, of the four nucleotides (C, G, A, U) of the messenger RNA (mRNA) molecules that act as an intermediary in the translation process. There are 64 permutations of bases, so the code has some redundancy: most amino acids are represented by two or more triplets (called codons) in mRNA.
The cracking of the genetic code began in 1961 when biochemists Marshall Nirenberg and Heinrich Matthaei at the US National Institutes of Health in Maryland found the first correspondence between a particular codon (UUU) and an amino acid (phenylalanine). By mid-1965 Nirenberg’s group had decoded 54 of the 64 possible codons; the last one (UGA) was finally decrypted in 1967, proving to be one of the three codons that, rather than encoding an amino acid, tells the translation machinery of the ribosome to stop.
At face value the genetic code looks arbitrary: it needn’t matter which codon corresponds to which amino acid, so long as the translation process can distinguish them. But is there really some chemical reason why a particular codon encodes a particular amino acid? That question has tantalised biochemists ever since the code was cracked. Even in the late 1960s researchers noted that there seems to be a relationship between the bases of a codon and the nature of its amino acid:2 the identity of the second letter of a codon seems to correlate with how hydrophobic its amino acid is, for example. This suggests that, when the genetic code arose – which must have happened around the onset of life itself, for the code is the same in all organisms – chemistry dictated the choices. But how?
Form and functionalisation
A common view of life’s origin is that the first proto-organisms contained neither proteins nor DNA, but only their intermediary RNA. For RNA is able both to serve as an information carrier – a nascent genetic material – and as a pre-enzymatic chemical catalyst. Eventually the genetic role was usurped by the more chemically stable DNA, while more versatile proteins took on the role of catalysis. Some believe that the latter handover came about as RNA molecules were modified by the attachment of short peptide chains, much as some protein enzymes now are functionalised with sugars or other non-peptide appendages. This would have turned the so-called RNA World into a Peptidated RNA World.3 Gradually, these peptide side-chains on RNA took over the catalytic responsibilities, and the job of RNA was simply to help assemble them.
If that’s so, it seems reasonable to suspect that the peptide-RNA association now preserved in the genetic code might reflect some common aspect of how the two types of monomer units in these molecules – amino acids and bases – were synthesised in the original protocells. This is what biochemist Stuart Harrison, working with Nick Lane of University College London, UK, and others, have now proposed.4 They show that the first letter of a codon correlates with the sequence in which the amino acids form via the biosynthetic fixation of carbon dioxide. That process of carbon fixation, using CO2 and H2 as the raw materials, occurs in much the same way in all organisms: it is the ‘core metabolic pathway’ of life, from which the Krebs cycle of aerobic metabolism emerged. Most amino acids are biosynthetic side-products of this core metabolism, and Lane and colleagues show that amino acids encoded by G at the first codon position are the first to be spun off from CO2 fixation. Amino acids encoded by codons starting with A are mainly the next to be produced, followed by those encoded by C-first codons and finally U-first.
In other words, the researchers say, amino acids encoded by purine (A, G)-first codons were probably the first to be recruited to the genetic code, for reasons to do with their ease of synthesis in metabolic pathways. As well as confirming the correlations with amino acid hydrophobicity for the second codon base, the team says that, at least for non-redundant codons, the third base correlates with the sheer size of its amino acid.
This suggests that random RNA sequences in the first cells will have interacted in a non-random way with amino acids – and we might then expect those sequences to be associated with the same amino acids in peptide and protein synthesis today. There may, in other words, be clues about the chemistry of the first organisms frozen into the genetic code.
References
1 M Cobb, Life’s Greatest Secret: The Race To Crack the Genetic Code. Profile, 2015
2 L E Orgel, Journal of Molecular Biology, 1968, 38, 381 (DOI: 10.1016/0022-2836(68)90393-8)
3 J T-F Wong et al, Life, 2016, 6, 12 (DOI: 10.3390/life6010012)
4 S A Harrison et al, BBA – Bioenergetics, 2022, 1863, 148597 (DOI: 10.1016/j.bbabio.2022.148597)










No comments yet