US scientists at the University of California, Santa Cruz, have made a key step towards nanopore protein sequencing, thanks to an ‘unfoldase’ enzyme. Mark Akeson’s team exploited this enzyme to unravel proteins and pull them through nanopores.
Though DNA sequencing techniques relying on similar nanopore-based processes are emerging that read longer fragments than predecessor technologies, there’s no equivalent yet for proteins. And while sequencers remain remote, the researchers see similar potential in their findings. ‘Protein translocation control for bench-top sequencing applications will require technical improvements,’ they write. ‘This is worthwhile because an optimised nanopore device could provide long reads of individual native protein strands.’
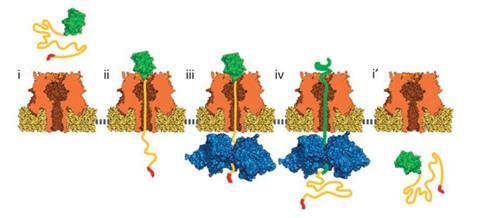
While important for biological function, proteins’ secondary and tertiary structures must be unfolded to pass them through a nanopore for any attempt at sequencing. In Escherichia coli the enzyme ClpX draws proteins through a nanopore into a chamber where they’re degraded. Akeson’s team realised it applies enough force to proteins to break down stable folds, and pull a protein through a pore fast enough to potentially be used in sequencing. So they put ClpX on one side of a lipid bilayer containing an a-haemolysin nanopore and put folded proteins with a trailing peptide tail on the other. They put a sequence known to target ClpX on the end of the tail. The peptide tail also contained a long sequence of negatively charged amino acids, which draws the protein to the pore when they applied a current across it.
When the team applied a current, ions flowed through the nanopore and they saw the ionic current vary as a protein passed through. The changes showed that the protein moved to the pore, with the tag poking through to the other side. ClpX bound the tag and moved along the chain, forcing the folded protein against the nanopore. That force and the pore’s electric field unfold the protein, allowing ClpX to pull the whole chain through. The team hope that these changes in the current could ultimately identify amino acid sequences.
‘This unusually strong work has considerable potential for use in biotechnology,’ says Hagan Bayley, at the University of Oxford, UK, who founded nanopore DNA sequencing company Oxford Nanopore. However, he calls using the approach for protein sequence determination ‘a distant dream’. ‘There may be more immediate applications such as analysing polypeptides produced by alternative splicing, an important source of protein variation,’ he adds.





No comments yet