Microfluidic technology is finally ready for forensic DNA profiling labs, as Rajendrani Mukhopadhyay reports
Microfluidic technology is finally ready for forensic DNA profiling labs, as Rajendrani Mukhopadhyay reports
TV forensics shows make DNA analysis seem so easy. Stick a sample in a machine, press a button, and get a DNA profile and a 3D image of a wanted criminal. Real-life DNA analysis is far more arduous, with many steps and several different instruments. But now microfluidics technology, where tiny volumes of fluids travel around miniature cartridges, is almost ready to deliver instant DNA readouts in true CSI fashion.
For over a decade, microfluidics has been touted as a way to bring together all the finicky steps of DNA analysis into one platform that can rapidly produce a person’s DNA profile. Now four commercial microfluidics developers - US companies IntegenX, NetBio, and Lockheed Martin in collaboration with ZyGEM; and the UK Forensic Science Service (FSS) together with the University of Arizona, US - say they are now getting ready to roll out prototypes to select customers in the next 12 to 18 months.
The technology has been a long time coming. For example, the US National Institute of Justice has funded research into microfluidics for forensic DNA analysis since the late 1990s, says Debra Figarelli, the DNA technical services manager at the National Forensic Science Technology Center (NFSTC) in Florida. And there is still some way to go. The prototype instruments will only be able to handle samples containing copious amounts of relatively pure DNA. Instruments to analyse samples taken from crime scenes will take at least five more years of development, say Figarelli and other forensic DNA experts.
Time wasting
Conventional DNA profiling was established in the 1990s. Now, thanks to TV shows such as CSI, juries expect to see DNA profiles as evidence at trials, whether or not it is needed, says Howard Goldstein, executive vice president of California-based IntegenX.
But conventional forensic DNA analysis takes time and technical expertise. There are three or four different steps that a sample has to go through before any data are generated, each step taking at least 30 minutes to run. And because a minimum of three instruments are involved, time is wasted taking samples from one to another.
The first step involves bursting open the cells to release DNA. If the sample came from a crime scene, like a blood stain on a bed sheet, the DNA is quantified by real-time PCR to get an idea of how much is available for analysis.
Next comes multiplex PCR to look at the short tandem repeats (STRs) in the DNA, which are unique to individuals. In STRs, two or more nucleotides (DNA building blocks) are repeated, with the repeated sequences next to each other. Forensic analysis surveys STR profiles with 4- or 5-repeated nucleotides that are stored in DNA databanks. The American database, run by the Federal Bureau of Investigation, is the National DNA Index System. The British database is the UK National DNA Database (NDNAD).
During the multiplex PCR, up to 15 STRs and a sex typing marker are analysed, explains Peter Vallone, a DNA analysis expert at the US National Institute of Standards and Technology, Maryland. The process takes about three hours. Once the PCR products are generated, they are separated based on size, usually by capillary electrophoresis. Once the DNA fingerprint is generated, it still has to be analysed.
In theory a skilled technician, with the help of robotics, can process multiple DNA samples in a day. However, in a forensic crime laboratory setting, samples have to follow a chain of custody so that technicians never lose sight of them for fear of contamination or mix-ups. Because of the time and labour investment, forensic crime laboratories commonly accumulate long backlogs of samples.
No more suck and squirt

Here’s where the appeal of microfluidics lies: it integrates processes and saves time. Over the past decade, the technology has matured to the point where it can handle several disparate chemical and biological analyses on a single chip or cartridge. No time is wasted transferring samples from one instrument to another. ’You free up a technician from all that suck and squirt,’ says James Landers, a microfluidics expert at the University of Virginia in Charlottesville and the chief scientific officer of ZyGEM.
Microfluidics channels nano- to picolitre volumes around a miniature cartridge. With such tiny volumes on a small footprint, processes are far more rapid. Susan Greenspoon, a forensic molecular biologist at the Virginia Department of Forensic Science, explains: ’With the advent and application of a microfluidic system, we have a real potential of consuming just a small fraction of what we consume now of our original casework samples by capturing, amplifying and profiling the DNA more efficiently.’
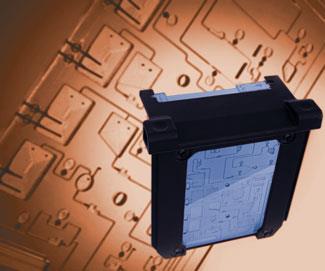

The time savings come in almost every step of the microfluidic process. Because researchers pull out just the right amount of DNA necessary for PCR, they don’t waste time going through a dilution step that takes place after extraction in the conventional analysis of reference samples. Then, for the PCR step, instead of using the block thermocycler traditionally used to cycle between the various temperatures, researchers like Landers use infrared heating, which is much faster. Microfluidics accomplishes electrophoretic separation in about five minutes, compared with 40 minutes for the conventional technology.
The developers of the new microfluidic platforms claim that their systems can process cheek swabs within two to three hours. But they are gunning for shorter timeframes of 45 minutes or less.
Keep it simple
Microfluidics also lends itself well to automation, so researchers can make platforms that are very simple to use. All a person potentially has to do is take a sample, stick it into a machine, press the ’go’ button, and wait for the instrument to spit out a DNA fingerprint. Andrew Hopwood at the FSS, Frederic Zenhausern at the University of Arizona, and colleagues recently demonstrated an instrument that delivers an individual’s DNA fingerprint within a few hours of taking a cheek swab.1 They hope that the instrument will be used by UK law enforcement officials to check if a person in custody can be matched to an unsolved crime using the National DNA Database.
In developing microfluidic platforms for DNA profiling, experts emphasise that they are not creating novel chemistry. For DNA profiles to hold up in a court of law, it is in the developers’ interests to stick to the same kind of chemistry for DNA extraction, amplification, and separation that has been long validated for the conventional technology. ’In many ways it is the same old chemistry but in a miniaturised form that now seamlessly integrates all three or four of the processes onto one device and closes it off so that it is not susceptible to contamination,’ says Landers.
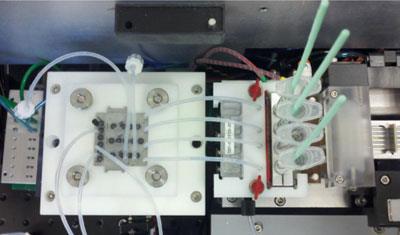

Cost has been a dominant factor in the development. Currently, a DNA profile on an individual costs about $100 (?62). ’If you offer [forensic scientists] a solution that is integrated into a single instrument that is 10 times faster but you tell them it costs $1000 a sample, it’s a no-go,’ says Landers. So plastics are very appealing as materials from which to fabricate the platforms. There are many to choose from that lend themselves well to affordable and simple mass production. Landers and others say that finding the correct plastics that permit optimal performance of the different chemistries has involved many trips back to the drawing board. But they are increasingly certain that plastics such as poly(dimethylsiloxane), polyacrylic, and cyclic olefin copolymer will play a dominant role.
The complete package
Because researchers want the platforms to be self-contained, they are embedding all the necessary reagents for the different analytical steps on the platform. This self-containment means that human hands are taken out of the picture and reduces the potential for contamination, a crucial issue in forensic analysis.
Incorporating the reagents onto the microfluidic platform involved some painstaking sorting, as Richard Selden, executive chairman of NetBio in Massachusetts, explains. He and his team devoted some time figuring how to store each type of reagent. ’There are two categories of reagents. One category is reagents that aren’t very stable for a long time, such as amplification enzymes. For those, we freeze-dry them. The second category is reagents that are stable for a year or more in liquid form such as simple buffers.’ Selden is now optimistic that their reagent-embedded microfluidic platform will be shelf-stable for at least a year.
Kinks remain in the technology, such as sensitivity. Researchers have to prove that the microfluidic platforms can handle casework samples with as much detection sensitivity as current techniques. ’When you are working with pristine, known reference samples, you have plenty of DNA and sensitivity is not an issue. But when you are working with evidence samples collected from a crime scene, you could have low levels of DNA,’ explains Figarelli. The microfluidic platforms have struggled to come up with the same level of sensitivity as the current technology used in forensic laboratories with crime-scene samples, she says. Quantitation of the amount of DNA in casework samples is also a critical factor where microfluidic technology has to catch up with the conventional technology.
Portable power

Because microfluidics integrates processes, the analysis becomes easier and more accessible to people lacking technical expertise. And because the integration into a microfluidic platform results in an instrument roughly the size of a computer tower, the analysis becomes portable. The portability and ease-of-use extends the applications of DNA fingerprinting beyond forensic crime-solving.
The instruments could be used to confirm the identities of individuals at immigration checkpoints, mass disaster sites, and military barracks. ’You can imagine you have someone at a border who claims the person accompanying them is their brother or daughter. You can test that claim,’ says Vallone. In a military setting, soldiers could test individuals coming into a base to confirm their identity.
Experts say the US Departments of Homeland Security (DHS) and Defense (DoD) are quicker and more aggressive in adopting the microfluidic technology than forensic crime laboratories. ’They are going to be looking at DNA analysis to get information, not to solve a crime,’ says Kevin Lothridge, chief executive of NFSTC. ’As soon as there is a system that’s operational and not problematic, they will deploy it for their specific forensic intelligence applications.’ Lothridge and Richard Mathies, a microfluidics expert at the University of California Berkeley, US, say that forensic crime laboratories are more cautious in adopting new technology because their experts need to be absolutely sure that the technology produces data that can be held up in a court of law.
Some experts mention processing casework samples at a crime scene but others are sceptical. ’I think it’s silly,’ says Bruce McCord, a forensic DNA expert at Florida International University, US. ’Laboratory people don’t want to be running samples at crime scenes because it might not be an appropriate place to be performing sterile laboratory procedures.’
Mini makeover
The manufacturers give a timeline for 12-18 months over which they will optimise their systems for small-to-mid-scale commercial manufacturing and provide instruments to customers. They are starting off with platforms that can handle up to 15 samples at a time. But because forensic laboratories will need to make investment in these systems worthwhile, ’over time, there will be need for higher throughput versions,’ says Selden.
Manufacturers are certain that they are onto something. Goldstein describes a time in October 2010 when he and his colleagues took IntegenX’s instrument to a forensics conference in San Antonio, Texas. ’We put it in a hotel room and we ran 45 profiles for people we invited to come and see it. The field team did it, not the science team. That alone - the instrument run in a hotel room by people who have minimal training - was exciting enough for the forensics folks.’
Developers of the microfluidic platforms say they are introducing a paradigm shift for forensic DNA fingerprinting by making it more accessible for a variety of applications. ’I like to think we’re democratising forensic analysis,’ says Goldstein. ’We have minimised the interaction with the science so the instrument can be used by anyone with minimal training - it is as easy to use as a toaster.’
It remains to be determined whether microfluidic platforms will convince forensic DNA analysts to punt on the conventional technology. A key step, explains Zenhausern, is going to be validation of the microfluidic measurements based on current standards in the industry. ’We have to demonstrate that the outcome of the measurement is exactly the same as the one done by the conventional techniques.’ Ensuring that the technology is robust and reliable in providing reproducible data in each forensic crime laboratory will take time. The time commitment is tricky for laboratories already struggling with backlogs. But developers of the instruments are hoping that the time invested will pay off handsomely in the long run.
Microfluidics certainly seems poised to broaden the applications of DNA fingerprinting. It will introduce new uses in identifying people for defence and security purposes. Time will tell whether it will give forensics laboratories a makeover to rival DNA analysis as seen on TV.
Rajendrani Mukhopadhyay is a freelance science journalist based in Washington DC, US
References
1 A J Hopwood et al, Anal. Chem., 2010, 82, 6991 (DOI: 10.1021/ac101355r)






No comments yet