A series of mistakes over 3.7 billion years has left us with a spectacular array of protein structures and functions, which are responsible for life itself, writes Bea Perks
A series of mistakes over 3.7 billion years has left us with a spectacular array of protein structures and functions, which are responsible for life itself, writes Bea Perks
Evolution tends to be thought of at the level of species - from the evolution of flowering plants to the ’ascent’ of man. Charles Darwin kicked it all off 150 years ago with the publication of On the origin of species. Darwin’s theory hasn’t changed, but what we now know, from research grounded on the foundations he laid, has dramatically changed the level at which we can think about evolution. That’s not to say that studying evolution at the species level is reserved for the history books, but there is a relatively new and steadily growing approach to evolution: evolution at the molecular level.
At a very basic level, weunderstand that evolution follows the accumulation of beneficial mutations in a gene sequence. Most of the time, gene mutations are detrimental. These are quickly removed by natural selection. But just occasionally, a mutation might improve (or at least not harm) an organism’s chance of survival. When that happens, the mutation will pass to the next generation. Evolution, over hundreds, thousands, millions and billions of years, has generated life as we know it. But while there is much talk of evolution on the macro scale - from the evolution of whole species to that of eyes or opposable thumbs - there has been relatively little mention of the molecular evolution that has underpinned these observable changes.
Protein evolution
On top of what we already know about evolution, researchers now have an eye-watering volume of gene and protein sequence data to hand. It is therefore becoming possible, using that data, to trace the evolution of proteins through time. Of course, sequence data alone cannot yield insight into the functions and structures of ancient proteins, or the processes by which their descendants evolved. But in the latest approach to the problem, scientists have begun to reconstruct ancient proteins and trace how they are most likely to have changed throughout evolutionary history to produce today’s proteins.
Joseph Thornton, an evolutionary and molecular biologist at the University of Oregon, US, and his team have shown how evolution tinkered with early proteins, and leaves the impression that complexity has evolved many times over.
Human and other animal cells contain thousands of proteins with functions so diverse and complex that it can be difficult to see how they might have evolved from a few ancestral proteins, says Thornton. His team recently studied the large family of nuclear receptors,1 which regulate key biological processes in animals by binding to specific DNA sequences and triggering the expression of nearby target genes. Nuclear receptors respond to hormones, nutrients and other chemical signals to regulate development, reproduction, metabolism and cancer.
There are some nuclear receptors that do not have to be activated by a chemical signal: they can trigger gene expression on their own. The received wisdom has held that the ancestral protein of today’s nuclear receptors would have been of this simpler type, implying that the complex capacity to bind and be regulated by chemical signals evolved independently in many lineages.
Using a database of the molecular sequences, functions, and atomic structures of hundreds of modern-day nuclear receptor proteins - from those found in sponges to those found in man - Thorton’s team reconstructed the biochemical characteristics of the ancestral nuclear receptor, which would have existed before the last common ancestor of all animals on earth - as much as a billion years ago.
The missing link
Finding an ’early’ nuclear receptor was tricky, because nuclear receptors are not found in all organisms. They are not found in plants or fungi; and nearly all the animal species that are known to have nuclear receptors, and whose genomes have been sequenced, have rather diverse selections of receptors - so they provide little information about which receptor genes were the first to evolve and which appeared later.

Luckily for Thornton, the genome of the sponge Amphimedon queenslandica was also published in 2010. Thornton’s team found that the A. queenslandica genome contains only two nuclear receptors (as opposed to the diverse selection in previously sequenced genomes), which they called AqNR1 and AqNR2.
His team also identified nuclear receptors in the genomes of two other recently sequenced species from the relatively distant evolutionary past: a curious flat, tiny (1mm diameter) animal called Trichoplax adhaerens and the sea anemone Nematostella vectensis, which contain four and 17 nuclear receptors respectively. Thornton concludes that, given the limited nuclear receptor diversification in these three separate species and knowing where each of them appeared in evolutionary history, these animals can shed light on early nuclear receptor evolution.
By reconstructing the family tree of nuclear receptor genes, Thornton found that AqNR1 represents the anciently diverged sister lineage to virtually all other nuclear receptors. Using a battery of biochemical and molecular assays, Thornton’s group showed that AqNR1 requires activation by a fatty acid. They also used computational methods to predict the three-dimensional atomic structure of the sponge proteins. This showed that they bound the fatty acid in a cavity very similar to that found in some receptors in mammals.
Thornton then worked his way backwards down the gene family phylogeny, computationally tracing features of the architecture and functions of nuclear receptor proteins back through their common ancestors, all the way to the progenitor of the entire receptor family, which existed in the very first animal perhaps a billion years ago. He found, contrary to current received wisdom, that the ancestral receptor required activation by a chemical signal - probably a fatty acid.
The underlying atomic mechanisms that allowed the ancestral protein to be activated by chemical signals are conserved in virtually all present-day descendants. In other words, it wasn’t a complex capacity that had to be evolved in many lineages further down the evolutionary line, but one that has been lost by a few. By analysing the structures of diverse nuclear receptors and comparing them to the ancestral template, Thornton’s team traced in detail how evolution tweaked the ancestral structure over time to yield new functions.
They found that in many receptor lineages, a few mutations subtly changed the size and shape of the cavity where the signalling compound binds, causing receptors to evolve partnerships with hormones or other new signals.
Other members of the receptor family became independent of chemical signals; these proteins, like switches stuck in the ’on’ position, evolved when simple mutations increased the intrinsic stability of the protein’s active conformation, removing the need for a chemical signal to activate gene expression.
Ancestral complexity
’If you just compare the receptors in modern humans, the evolutionary events by which they could have evolved are not obvious. It may look as if the complex functions of each protein evolved independently,’ says Thornton. ’But when we traced these proteins from their ancestor through time, we saw how evolution tinkered with the ancestral form, producing an incredible diversity of protein functions and the ability to interact with many different chemical signals.’
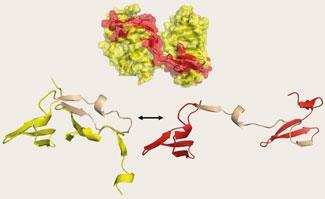
Nuclear receptors are a ’great case study in protein evolution’, says Thornton, adding that it is likely that other protein families, when studied in similar detail, will turn out to have diversified by a similar kind of tinkering. ’We predict that, when sufficient data are gathered to allow detailed evolutionary reconstructions, it will become apparent that most protein superfamilies diversified by subtle modification and partial degradation of ancient, deeply homologous functions. Invoking the evolution of wholesale ’’novelty’’ will seldom be necessary.’1
That’s all very interesting, of course, but is it really useful? Very much so, Thornton explains: ’Structure-function relationships are the fundamental object of knowledge in protein chemistry; they allow us to rationally design drugs, engineer proteins with new functions, and understand why mutations cause disease.’ Unfortunately, revealing how the sequence of a protein determines its function using traditional laboratory manipulations is difficult, because the number of possible sequence variations in any protein family is too great. ’The solution lies in studying evolution, which has been a massively parallel experiment in diversifying and optimising proteins, conducted over vast periods of time’ says Thornton.
One person who has found protein evolution both interesting and useful is Frances Arnold, a chemical engineer at California Institute of Technology (Caltech) in Pasadena, US. Arnold, together with Willem ’Pim’ Stemmer, chief executive of the Californian biopharmaceutical firm Amunix, has just been awarded the Charles Stark Draper prize - widely considered the equivalent of the Nobel prize for engineering in the US - for developing directed evolution: ’a method used worldwide for engineering novel enzymes and biocatalytic processes for pharmaceutical and chemical products.’
Inspired by the real-life evolution of proteins, Arnold developed a system that literally ’directs’ the evolution of proteins by repeated rounds of mutation and selection. ’Evolution happens by the accumulation of beneficial mutations one at a time through functional proteins,’ says Arnold. ’And that’s a very simple algorithm; it doesn’t require that you screen large numbers of things if you can go through single mutational stops.’
Mimicking evolution
Arnold began to evolve proteins by randomly mutating the gene of a known protein, screening its mutant offspring a few-hundred or a few-thousand at a time, finding one beneficial mutation, and repeating the process starting from that mutant gene. ’That’s just mimicking how adaptive evolution happens,’ she says. ’It doesn’t fully mimic it, because we miss the entire neutral mutation pathways,’ she says, ’but people are starting to try to insert neutral mutations into directed evolution as well.’ Neutral mutations will be passed on from one generation to the next if they pose no threat to an organism’s survival. They shouldn’t be ignored because, further on down the line, they might help give rise to a beneficial mutation.
Another important lesson from adaptive, real-life, evolution has come from recognising which proteins and what functions are evolvable. ’We’ve learnt that the proteins that evolve readily in nature also evolve readily in the laboratory,’ says Arnold.
’For example, secondary metabolic pathway enzymes, like cytochrome P450s and other enzymes whose specificities and functions have varied all over the map tend to readily change in the laboratory,’ she says. ’And the converse is true, proteins whose functions have not shown great diversity in nature tend to be more difficult to engineer’
Arnold comes from what she calls an engineering and problem solving background. ’I was looking for the most effective means to engineer proteins, the sequence basis of whose functions we just don’t understand. So a rational design pathway to achieve very specific functions just wasn’t realistic, and evolution seemed to be a much more promising route.’
Just as Arnold keeps up with the world of natural protein evolution, particularly work from Thornton’s lab, protein evolutionists are now reading the directed evolution work. ’We have one huge advantage,’ says Arnold, ’and that’s data.’
Although directed evolution started from an engineering perspective, researchers from different backgrounds and interests have come to it specifically to answer questions about how proteins evolve. ’If you look at people like Joe Thornton or a number of people whose primary interest is in evolution, they are now using directed evolution to try to understand how proteins can adapt,’ she says.
The traditional view that proteins possess absolute functional specificity and a single, fixed structure conflicts with their marked ability to adapt and evolve new functions and structures, says Dan Tawfik of the Weizmann Institute of Science in Israel.
’Natural selection is yielding molecular machines with breathtaking performance,’ says Tawfik. ’New functions can evolve within years or even months, as happens naturally with things like drug resistance.’
Tawfik’s team argues that proteins exhibit ’functional promiscuity’, which is an essential component of a protein’s evolvability. He correlates promiscuity with conformational diversity. In antibodies, for example, increased affinity for a ligand - clearly an essential characteristic of an antibody - leads to decreased binding-site flexibility. Conversely the P450s, as mentioned by Arnold, demonstrate increased promiscuity - relatively broad substrate scope - with increasing flexibility.
But is it really chemistry?
The study of protein evolution lies at the interface of chemistry and biology, and relies to a large extent on engineering. Each discipline can claim it as their own, but it is often dismissed as ’not really biology’ or ’not really chemistry’.
It’s definitely chemistry, says Arnold the engineer, who nevertheless files directed evolution firmly under ’synthetic biology’. Apart from anything else, scientists have a habit for winning Nobel prizes in chemistry for work on proteins.
Back in 1946, the Nobel prize in chemistry went to James Sumner of Cornell University, US, who won half the prize ’for his discovery that enzymes can be crystallised’; and John Northrop together with Wendell Stanley, both of the Rockefeller Institute for Medical Research in Princeton, US, shared the other half ’for their preparation of enzymes and virus proteins in a pure form’.
In 1972, half the chemistry Nobel prize went to Christian Anfinsen of the US National Institutes of Health with the other half shared by Stanford Moore and William Stein, both from Rockefeller University in New York, US, for fundamental work in protein chemistry. Anfinsen had shown that the information for a protein assuming a specific three-dimensional structure is inherent in its amino acid sequence, which was the starting point for studies of protein folding. Moore and Stein received the prize for discovering anomalous properties of functional groups in an enzyme’s active site, as a result of the protein fold.

In 2009, the prize went to Venkatraman Ramakrishnan, Thomas Steitz and Ada Yonath ’for studies of the structure and function of the ribosome’. In 2008 it was awarded jointly to Osamu Shimomura, Martin Chalfie and Roger Tsien ’for the discovery and development of the green fluorescent protein, GFP’.
On Frances Arnold’s website,3 team members have contributed to a ’How would you explain synthetic biology to Darwin?’ section.
In Arnold’s words: ’You had it just right! And now I can build new components of life - for example, the proteins of which we are made - by artificial selection. When I decide who gets to reproduce, these components evolve just like the organisms that make them do, because they are encoded by the same stuff and follow the same principles
of evolution that you showed us for all forms of life.’
References
1 J T Bridgham et al , PLoS Biol., 2010, 8, e1000497, (DOI: 10.1371/journal.pbio.1000497)
2 N Tokuriki and D S Tawfik, 2009, Science, 324, 203






No comments yet