Sensors with molecular probes that fold around their targets, in combination with electrochemical detection, are being heralded as the ultimate user-friendly 'just add sample' device. Michael Gross reports.
Sensors with molecular probes that fold around their targets, in combination with electrochemical detection, are being heralded as the ultimate user-friendly ’just add sample’ device. Michael Gross reports.
In the beginning, there was the folding problem. Many research groups around the world study how proteins and nucleic acids fold, sometimes to get to grips with the biological systems, but often just for the intellectual challenge of working out how these long chain molecules fold up, with remarkable speed and efficiency, into highly specific 3D structures.
Kevin Plaxco had spent several years researching folding when he set up his own research group at the University of California, Santa Barbara (UCSB), US in 1999. He decided to do something practical with his folding experience and hit upon the idea that folding could solve a long-standing problem in the area of biosensors. ’By engineering biomolecules such that they fold only when bound to their targets, I wanted to couple target recognition with an enormous change in the physics of the biomolecule and use this change, in turn, to trigger an electronic response,’ he explains.
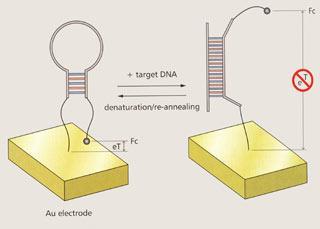
Stretching out sensors
Plaxco’s first device was a gene sensor dubbed E-DNA, containing a folded-up probe sequence of DNA which stretches out when it binds the exactly matching DNA target sequence. This stretching allows the DNA to move a ferrocene label fixed at one end away from the gold surface to which its other end is immobilised. This suppresses the electrochemical signal produced by the label (Chemistry in Britain, September 2003, p20).
As DNA sensors go, E-DNA appears promising in that it is sensitive, sequence-specific, reagentless and fully electronic (and thus small - the sensor can fit on your palm). But DNA can be used for many different things, including molecular recognition of other kinds of targets.
DNA molecules with highly specific binding properties selected from a large pool of random sequences are known as aptamers. Since the discovery of this concept in 1990, aptamers with many different specificities have been reported and used in many applications including fluorescence-based sensors, known as aptamer beacons. Last year, Plaxco’s group, in collaboration with their neighbour at UCSB, Nobel laureate Alan Heeger, used one such aptamer, with a specificity against the blood coagulation factor thrombin, to convert the E-DNA into an electronic aptamer-based (E-AB) sensor that could detect as little as 20 nanomolar concentrations of this protein directly in blood serum.
Sensing dirty blood
The high specificity of this first E-AB sensor stems from the fact that the signal change could only be triggered by the DNA aptamer folding around the thrombin target, an event which is much more specific than simple adhesion processes and allows the sensor to work in blood and other grossly contaminated materials. Like the E-DNA this sensor had the advantage of being reagentless and fully electronic, implying that the technology could be developed into a simple, hand-held device, which the end-user would only have to immerse into the sample to be analysed.
’Making molecular beacons electronic has a huge impact on their convenience,’ Plaxco told Chemistry World. ’And because the electrochemical background of a typical real-world sample like blood serum is many orders of magnitude lower than its fluorescence background, electrochemical sensors work in even the dirtiest samples.’
But both the thrombin sensor and the original E-DNA sensors had the disadvantage of being ’signal-off’ architectures - the signal recorded in the absence of the target is weakened when target molecules are detected. Another group working in this area, led by Ciara O’Sullivan at the Universitat Rovira i Virgili at Tarragona, Spain, could go one better. They also developed an aptamer-based sensor for thrombin, but due to subtle chemical differences in the design of the aptamer sequence, their device works in the more desirable signal-on mode.
Plaxco’s group, in contrast, had to invest additional work into its E-AB sensor in order to convert it to signal-on mode, which led to a signifiant improvement in sensitivity. ’By making our sensor signal-on,’ Plaxco explains, ’we go from a 30 per cent signal loss at saturating target to a 300 per cent signal increase, which improves our sensitivity by an order of magnitude.’
However, O’Sullivan adds words of caution: ’Aptamers are inherently highly sensitive to the presence or absence of cations, which may seriously affect the assay sensitivity - in either a positive or negative manner. In the case of the thrombin aptamer, our experiments have shown that this aptamer can as easily be exploited to detect potassium as to detect thrombin.’
Illicit drugs
Recently, the Santa Barbara group also developed a highly sensitive device to detect cocaine in blood serum and in the presence of the masking and cutting agents often employed in the illicit drug world (Chemistry World, April 2006, p 25). Their latest development is a sensor for the blood protein PDGF, thought to be diagnostic for kidney and other cancers. Unlike the fluorescence-based sensors that other groups have developed for the same purpose, this electrochemical device can be used in blood serum, which only needs to be diluted two-fold with water or buffer to reduce the sample’s ionic strength. (At very high ionic strength, the aptamer tends to collapse, which leads to a high background current and reduced sensitivity.)
Commercial applications for these sensors might not be that far away. ’What makes this work so interesting is the straightforward and rapid way it could impact healthcare and industry outside of the laboratory,’ says Faiz Kayyem, who developed one of the first useful electrochemical sensors, founded the company Clinical Microsensors, which he sold to Motorola, before moving into venture capital at Efficacy Capital, San Diego, US.

’If the aptamer approach turns out to be generalisable and perhaps a bit more sensitive, then one can easily see the potential of this work to move testing out of the central lab and into the places where it’s needed: the home for patients monitoring their therapuetic drug and enzyme levels; the field for law enforcement and border personnel charged with detecting illicit drugs; the farm and food packer for food safety monitoring, for example, all with a device as small, inexpensive to manufacture and easy to use as a diabetic’s glucose sensor.’
Michael Gross is a science writer based in Oxford, UK.
Further Reading
- Y Xiao et al, Angew. Chem. Int. Ed. 2005, 44, 5456.
- A -E Radi et al, J. Am. Chem. Soc. 2006, 128, 117.
- Y Xiao et al, J. Am. Chem. Soc. 2005, 127, 17990
- B R Baker et al, J. Am. Chem. Soc. 2006, 128, 3138






No comments yet