Sarah Houlton talks to Hagan Bayley, the 2008 Chemistry World Entrepreneur of the year
Hagan Bayley first became interested in membrane proteins during his PhD. Now, more than 30 years on, he’s using his knowledge about these proteins to make nanopores that can be used to sequence DNA. And the company set up to commercialise the technology, Oxford Nanopore, is showing a lot of promise - it has managed to secure a significant amount of funding even in the current challenging economic climate.

Brought up in north Wales, Bayley studied chemistry at Oxford, before moving to Harvard for his PhD as his supervisor, Jeremy Knowles, joined the academic staff there. ’Membrane proteins were a new venture for Jeremy’s lab back in the 1970s, and no structures had been determined or sequences identified for any of these proteins,’ he says. ’We were interested in finding out how these proteins sit within biological membranes, and by making radioactive, photoactive substances that partitioned into the lipid bilayer, we could map the regions of the protein that had been labelled, and discover what was in the membrane.’
On finishing his PhD, he moved across town for a postdoc with Gobind Khorana at MIT. Khorana was working on another membrane protein, bacteriorhodopsin, which they’d just managed to sequence. It was difficult to sequence proteins back then, especially membrane proteins, but nevertheless it was still easier than sequencing genes. ’That opened up a huge number of possibilities for membrane biology, and it was an incredibly interesting time,’ he says.
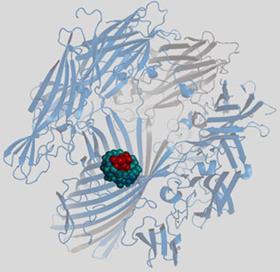
Next stop was an assistant professorship at Columbia University in New York, and six years later a move to the Worcester Foundation in Massachusetts enabled him to concentrate on research. And it was here that he first became interested in α-haemolysin, which is at the heart of Oxford Nanopore’s technology.
Bayley moved to Texas A&M University in 1996, and by this time he was working with Eric Gouaux, who had solved the crystal structure of α-haemolysin. ’This allowed us to forge ahead with the protein engineering that eventually led to our interest in nanopore sensing,’ he explains. He returned to the UK and a chair at Oxford University in 2003, and within a couple of years Oxford Nanopore was born.
$1000 genome project
’There were three of us at the start,’ he explains. ’I was the scientific founder; Gordon Sanghera, who had worked on glucose sensing with Allen Hill at Oxford and then at Medisense and Abbott, was the most entrepreneurial one; and Spike Willcocks, who had studied chemistry at Oxford and moved into finance, was keen to get closer to the science again. Initially, the company was set up to explore the possibilities of sensing single molecules, but we realised that the technology had enormous potential in DNA sequencing.’
Bayley’s lab had already been given an academic grant as part of the US National Institutes of Health’s $1000 genome project, and in 2006 published a paper that first showed that the nanopore technology could be used to identify individual bases from DNA. ’It was a logical next step for the company to focus on,’ he says. ’It’s a very exciting area to be in - it’s high-risk but has the potential for a high pay-off if we succeed.’
There certainly seems to be optimism in the financial community that it will work. In 2006, they raised about £7.5 million, led by London-based IP Group. A further £10 million was raised in 2008, again in the City of London. And in January of this year, they raised a further £14 million. ’Altogether, we’ve raised about £32 million, despite the difficult environment,’ Bayley says. ’I think this is mainly because people are still tremendously excited about advances in DNA sequencing.’ About a third of the money came from the US company Illumina, whose sequencing enterprise started life as University of Cambridge spin-out Solexa.
NIH’s $1000 genome challenge set the goal of sequencing an entire human genome for $100,000 by 2010, and for $1000 by 2015. ’It’s always hard to get the real numbers, but I believe Illumina has already achieved the $100,000 genome well in advance of the target date, and maybe this year they will even get down to the $10,000 genome,’ he says. ’However, sequencing in this way still has underlying fixed costs; it is done on chips, and first the DNA has to be amplified, tagged with costly fluorescent reagents, and then very expensive charge-coupled device cameras are needed to record the data. We feel that nanopore sequencing should be a lot cheaper - it needs no reagents, and gives a direct readout of the sequence, with no need for expensive equipment.’
Binding events
Oxford Nanopore’s technology was originally developed for sensing. The nanopores could be created from an inorganic material such as silicon nitride by making a hole in it, but there are manufacturing issues with this. It is far more practical to exploit a nanopore that occurs naturally, and this is where the a -haemolysin comes in. ’It is secreted as a water soluble monomer by Staphylococcus aureus bacteria, and self-assembles in lipid bilayers, forming heptamers with a hole in the middle,’ Bayley explains. ’It’s a very simple protein, and we’ve been able to engineer it into many different things, including forms that can be used as a sensor. If you insert one of these assemblies into a biological membrane with a salt solution on each side, when you apply a potential to the system you can read the current that goes through the pore, and this varies according to what is occupying the pore (see Chemistry World, April 2009, p23).
’When something binds inside the pore, it perturbs the current, usually making it lower. So you can watch a substance binding to the pore and coming off again at a single molecule level, and different molecules block the current to different extents and for different periods of time.’ It’s also very sensitive - they found that even the two enantiomers of the painkiller ibuprofen block the channel to a different extent. ’So we thought that if we could do that it would actually be easier to distinguish the four bases of DNA.’
And that turned out to be the case. The aperture of the nanopore is a couple of nanometres in diameter, which is just wide enough for a single strand of DNA to enter, but not a double strand. ’There are two ways that this might be exploited to sequence DNA,’ Bayley says. ’One is to pull the single strand through the pore and identify the bases as they go past, and the other is to cut the bases off one at a time, have them drop into the pore, and identify them as they emerge on the other side. We’re working on both of these areas. We have already managed to show that we can identify the bases in the pore when they are separated, and also that it should be possible for DNA strands.’
The next challenge is to couple an enzyme to the top of the pore that will chop off the bases and deliver each and every one into the pore. They also need to make the whole system highly parallel. ’The systems that Illumina have produced will sequence millions of DNA strands simultaneously on a chip,’ he says. ’Nanopore sequencing is much faster - between one and 10 milliseconds per base - but I still think we will have to have a thousand, or even ten times that many, of these pores working in parallel.’
Sequencing uncut strands poses a further problem - there is a voltage threshold that needs to be met before the DNA will thread into the pore, but at that voltage the DNA passes through at speeds of around one microsecond per base. ’That’s far too fast for us to measure the current levels,’ Bayley admits. ’So we are trying to devise ways that will allow the DNA to go through the pore more slowly, more like a millisecond per base. We need it to be fast, but not so fast that we can’t read the bases.’ Slowing it down will involve either engineering the pore, or perhaps using an enzyme to thread the DNA in. ’You could imagine a polymerase that would make a replica strand and thread it in, a nuclease that will eat up the DNA but pull it through as it eats it up, or a helicase, an enzyme that will take double-stranded DNA and convert it into a single strand, pushing it through the pore in the process.’
Out of academia
Bayley says he’s a deep believer in fundamental science, but if that science has commercial possibilities, then an academic lab is not the best place to progress it. ’If there is a possibility for a commercial product, then it is appropriate to farm it out to a company,’ he says. ’Bodies like EPSRC [Engineering and physical sciences research council] are trying to push academic science more directly into the commercial world, and I think that is risky. I don’t think a cure for cancer, for example, will come from a Grand Challenge - Nixon tried that and it didn’t happen! It will come from basic research.’
But he does believe that spinning out a company to exploit new science has several positive effects. ’First, there’s the scientific feedback. Basic scientists are very often opportunistic - they see an interesting problem and work on it, but if it doesn’t work after, say, a year, they will give up. In industry you can’t do that - you have a specific goal and you have to execute it, which may well be difficult.
’Scientific funding is often piecemeal and time-consuming to obtain and manage. Setting up this company has allowed me to have five company-sponsored postdocs in my academic lab. When I first came back to the UK, I submitted a proposal on stochastic sensing. I’d got matching funding from the Ministry of Defence for £0.5 million, but the proposal was rejected by EPSRC. That’s when I decided the only answer was to set up a company, which has now been funded to the tune of £32 million, and some small fraction of that comes back to my academic lab to do basic research in the nanopore area.’
And he hopes that setting up the company will bear fruit in the fiercely competitive field of DNA sequencing. ’We’re trying to climb Mount Everest, not Snowdon - this is a really difficult task!’ Bayley says. ’We have a well-funded company, a large amount of know-how and 60 people working for us, most of whom are scientists. If anyone is going to get nanopore sequencing to work, it will be us.’
Sarah Houlton is a freelance science journalist based in London, UK











No comments yet