The way spider silk proteins can be stored as a fluid but spun instantly into fibres is all down to their end parts, research suggests
The way spider silk proteins can be stored as a fluid but spun instantly into fibres is all down to their end parts, European scientists have discovered. Two groups, working on opposite ends of the spidroin proteins that make up silk, have begun to piece together how changes in pH, salt balance and the mechanical force applied during spinning combine to control the aggregation of the proteins into super-strong silk fibres.
Spidroin proteins are mostly made up of a repeating sequence of amino acids, rich in alanine and glycine, which gives the fibres their exceptional mechanical properties. ’But we realised quite quickly that these core parts lack any controls for making fibres out of solution,’ explains Thomas Scheibel from the University of Bayreuth in Germany, who led one of the research teams.
So researchers turned their attention to the end parts or ’terminal domains’ of the proteins, since similar domains control assembly in other proteins like collagen. Various experiments have shown that these terminal domains affect fibre formation, says Scheibel, but exactly how this works remained a mystery.
The terminal domains at either end of spidroins have different effects. Scheibel teamed up with Horst Kessler’s group at the Technical University of Munich in Germany to study the carboxyl (C) terminus,1 and another team - led by Stefan Knight and Jan Johansson from the Swedish University of Agricultural Sciences in Uppsala - took on the amino (N) terminus.2
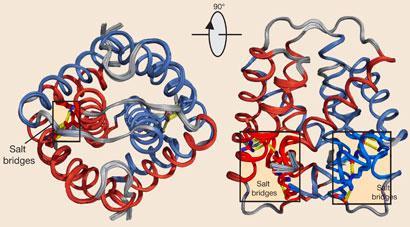
Kessler explains that the C-terminal domains pair up to form dimers. The protein chains are intertwined and joined by a disulfide bond, and the chains of repeating silk units stick out at an angle of 60? to each other. ’To form a thread, they have to go parallel,’ he says, ’which means the terminal domain has to unfold a little.’
To work out how this happened, the team looked at the solution structure of the terminal domains by nuclear magnetic resonance (NMR). They saw that as they switched the salt solution around the protein from sodium chloride to sodium phosphate - which is one of the things that happens as the silk moves towards a spider’s spinning duct - two salt bridges within the structure were disrupted, allowing the silk chains to align together and form fibres. Kessler adds that this partial unfolding is also affected by the shear stresses the proteins experience while flowing towards the spinning duct.
At the opposite end of the spidroin chains, the story is a little different. Knight and Johansson’s team found that the key trigger for the N-terminal domain’s aggregation is the drop in pH in the spider’s spinning apparatus. ’It’s a highly charged protein, with a positive and a negative pole,’ says Knight, adding that in the silk stores the domains pair up head-to-tail to form dimers. ’But there’s more negative than positive charge, and as the pH drops below 6.3, we think some of the acidic groups are protonated.’ This could result in a better charge balance or trigger some kind of conformational change in the protein and favour aggregation into larger structures like fibres.
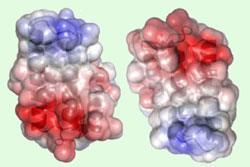
The Swedish group looked at the structure and amino acid sequence of the N-terminal domain, and tried to identify likely amino acids to be protonated. They tried replacing several of the most obvious candidates, but while some of these changes affected how the spidroins aggregated, they didn’t affect the pH of the transition. ’We still don’t fully understand that aspect of the switching mechanism,’ says Knight, but he adds that it does look like a single amino acid side chain is responsible.
The combined results of both groups provide a solid platform for studying spider silk at the molecular level, says Knight. ’It’s also important that the changes in conditions that have these effects are physiologically relevant and resemble what’s actually going on inside the spiders.’
Scheibel is excited about the implications of understanding what controls the assembly of spider silks. ’By understanding the contribution of these termini, we can design proteins and spin them into biomimetic fibres,’ he says. Incorporating certain motifs into those proteins would allow chemical or biological modifications of the silk fibres, Scheibel adds, ’For example we could add metal to get a conducting shell around the fibre, or immobilise enzymes or dyes on them.’
Phillip Broadwith
References
1. F Hagn et al, Nature, 2010 DOI: 10.1038/nature08936
2. G Askarieh et al, Nature, 2010 DOI: 10.1038/nature08962





No comments yet