Protein binding is not ruled by increases in entropy when ligands displace water, contrary to long-held 'hydrophobic effect' theory
US researchers have dealt a severe blow to the idea of a single ’hydrophobic effect’ that can help explain how all drugs dock with proteins. A team led by Harvard University’s George Whitesides has shown that water structures in binding pockets can cause hydrophobic interactions with thermodynamic behaviour differing from a widely used model. ’One conclusion from that is that there is not one hydrophobic effect but a distribution of hydrophobic effects,’ Whitesides tells Chemistry World.
Thermodynamically favourable associations between non-polar surfaces in aqueous solutions are often the most important contributor to how ligands like drugs or biomolecules bind with proteins. Research reaching back at least 50 years has built an understanding of this process where hydrophobic parts of similarly shaped ligands and binding sites meet. This ’lock and key’ analogy comes largely from studies partitioning non-polar molecules between hydrophobic liquids and water, which are thermodynamically dominated by entropic effects.
’There is still the pervading sense that the hydrophobic effect is entropically driven,’ says Phillip Snyder, a member of Whitesides’ group. ’The idea was that when the ligand associates with the protein, water was released,’ Whitesides explains. ’The entropy from that drove the interaction of these two hydrophobic surfaces.’
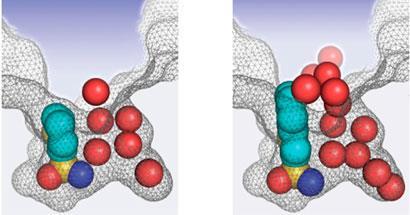
Whitesides, Snyder and colleagues studied hydrophobic binding effects in the enzyme human carbonic anhydrase II (HCA). They measured changes in enthalpy, entropy and Gibbs free energy occurring both when ligands bound to HCA, and when the same molecules were partitioned between octanol and water. The researchers also used x-ray crystallography and molecular modelling to study molecules’ locations in the ligand-enzyme complexes.
Entropy again played the largest role in hydrophobic effects in the octanol-water system. However, energy released from breaking down hydrogen bonding structures in the water around the enzyme and ligand was responsible for their binding. ’In this case it was enthalpy that we observed to be more important,’ Snyder says. ’What really drives the association is the release of those structured networks in the cavity and around the ligand,’ Whitesides adds.
Ulf Ryde, professor of theoretical chemistry at Lund University, Sweden, called the findings ’a small but carefully performed step towards understanding the hydrophobic effect’. ’The strength is the combination of several experimental and theoretical methods and also the careful and nuanced discussion of the data,’ he says. ’The weakness is that only a single protein and type of ligand are studied, so we don’t know how general the results are.’
Nevertheless, Snyder asserts that these results undermine hopes of finding easy yet accurate ways to predict binding in pharmaceutical research. ’Every drug, every protein is going to be different,’ he says. Whitesides hopes that studying protein-drug interactions in this way will still help make early stage drug discovery quicker than relying on existing models, though. ’We’ve had a model that grew out of lock and key that really hasn’t worked for a long time,’ he says. ’This is at least a different line of reasoning to pursue. With a bit of luck something will grow from it that’s more predictive and useful.’
Andy Extance
References
Proc. Natl. Acad. Sci. USA, 2011, DOI: 10.1073/pnas.1114107108






No comments yet