The molecule that revolutionised and illuminated cell biology started with a jellyfish. Lewis Brindley tells the story of this year's Nobel prize for chemistry
The molecule that revolutionised and illuminated cell biology started with a jellyfish. Lewis Brindley tells the story of this year’s Nobel prize for chemistry
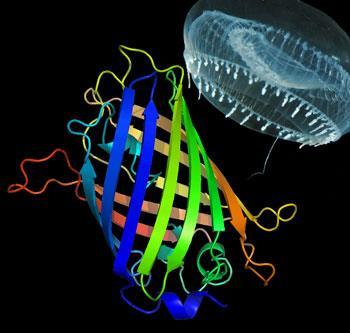
As Osamu Shimomura collected his one thousandth jellyfish of the day, on a hot summer afternoon in 1961, he had no idea that the painstaking work would eventually net him a Nobel prize. But Shimomura’s discovery of green fluorescent protein (GFP) later that year set in motion a series of developments that revolutionised the way we look inside cells.
The two men who share the Nobel prize in chemistry with him this year are Martin Chalfie and Roger Tsien. Each played a critical role in turning an unusual jellyfish protein into compounds that are routinely found in cell biology labs across the world and continue to provide new insights into diseases such as cancer, HIV and Alzheimer’s disease.
The great jellyfish hunt
Shimomura was born in Japan in 1928, and his early life was blighted by war. He was only 12 kilometres from the atomic bomb that struck Nagasaki - close enough to be temporarily blinded by the flash.
His determined pursuit of a scientific career was eventually rewarded with a PhD from Nagoya University for discovering the protein responsible for the luminescence of a sea-dwelling firefly. This work caught the eye of Frank Johnson at Princeton University in the US, who recruited him to study the green-glowing jellyfish Aequorea victoria in a project that would last more than 20 years.
Every summer, Shimomura would sail out into the bay of Friday Harbor, San Juan Island, Washington, when the sea became thick with jellyfish. During 1961, he extracted the faintly glowing rings from around 10,000 of the mouse-sized jellyfish and squashed them through a filter. After isolating a protein from the glowing ’squeezate’, Shimomura was surprised to find that it emitted blue light, not green. This, it turned out, was aequorin - a protein that produces blue light in the presence of calcium ions.
Shimomura realised that something else was converting this blue light into the green glow. Eventually he isolated a second protein, which was able to absorb both blue light and ultraviolet light and then fluoresce green. This protein was referred to as green fluorescent protein (GFP) - it was a simple name that stuck.
For the next 17 years Shimomura continued to harvest jellyfish to identify the chemical makeup of the glowing proteins. ’We needed a lot of material, so our schedule required us to catch around 50,000 jellyfish every summer,’ says Shimomura. ’So in total we probably collected over 850,000.’
From this great collection, Shimomura was finally able to characterise the chemical structure of the GFP chromophore in 1979 and lay the groundwork for future discoveries.

Seeing through it
In 1988, Martin Chalfie, a biological scientist from Columbia University in New York, attended a seminar during which GFP was briefly mentioned. Chalfie remembers being so excited by the idea of a fluorescent protein that he didn’t listen to any other speakers that day.
’The reason I was so excited was because I was working on a millimetre-long roundworm called C. elegans - which is transparent,’ says Chalfie. ’I realised that if I could express the fluorescent protein in the roundworm, I could watch cellular processes as they happened under the microscope.’
Chalfie needed to find the gene responsible for making GFP, and it didn’t take long to find someone who was working on it - a biologist named Douglas Prasher at Woods Hole Oceanographic Institution in Massachusetts.
Prasher was the first to envisage how GFP might function as a fluorescent tag. Being small enough to not hinder the action of other proteins and responsive to ultraviolet light, Prasher reasoned that GFP would be perfect for tracking cancer cells. By 1992, he had isolated the GFP gene and published a paper detailing the sequence of the 238 amino acids that make up the protein.
On hearing of Prasher’s results, Chalfie decided to instruct his graduate student, Ghia Euskirchen, to begin work putting the GFP gene into the genetic code of E. coli bacteria. Only one month later Euskirchen succeeded - producing E. coli cells that glowed brightly despite having none of the other jellyfish enzymes that were thought to be needed.
Many organisms fluoresce naturally, such as fireflies or tropical fish, but the majority of these processes involve other components that are used up during the luminescence. Chalfie found that GFP was unique - the protein itself was all that was needed to convert ultraviolet light into a bright shade of green.
In the genes
Chalfie says that luck played a role in winning the race to express the gene. ’At least three other teams had tried to express GFP before us, but their attempts all failed. We later found out that this was because they had used a slightly different process.’
Whereas Chalfie’s team amplified only the precise sequence, others cut appropriate sized fragments from Prasher’s clone. However, this left small sequence fragments attached to either end of the strand that interfered with the correct formation of the protein.
Despite the success he achieved, Chalfie was still surprised by the advances that GFP led to. ’I certainly realised that attaching fluorescent tags to a subset of cells could be important. But what I never could have guessed was all the wonderful adaptations and modifications that others have come up with, which really made GFP into a useful and effective tool.’
It wasn’t until Roger Tsien became involved in 1994 that explanations for GFP’s behaviour were revealed. Tsien was already well-known in the field for giving unprecedented views inside cells, for example, developing dyes that change colour in contact with calcium ions.
Jeremy Sanders, who supervised Tsien during his PhD at the University of Cambridge, UK, says the work Tsien did before his research on GFP ’opened up a whole new world for looking at what was happening inside living cells’. ’At some point during the 1980s, nearly a third of all papers published in the field of physiology cited Tsien’s work.’
To begin with, Tsien charted how the chromophore of GFP was formed in the 238 amino acid chain. The secret lies in the way the protein folds itself - wrapping up into a barrel-like shape, with three key amino acids (a serine-tyrosine-glycine sequence) folded tightly together in the centre. Tsien showed how these three amino acids could react together with ambient oxygen to create the fluorescent chromophore.
Next, he set to work making improvements. ’The wild type of GFP was made for the jellyfish’s own purposes, so it obviously wasn’t ideal for ours,’ says Tsien, noting that GFP tended to lose its ability to glow over time and green was not always ideal for practical studies.
’We found that a specific amino acid could change the primary colour of emission, but other amino acids also played smaller roles,’ he explains. By gradually making mutations in the gene to carefully change the amino acids, Tsien was able to tweak GFP to create a broad palette of colours and improve overall brightness.
The new colours were an important step forward, as they allowed scientists to image multiple different cells simultaneously and study how different systems interact in real time.
Over the brainbow
One thing Tsien had not yet managed to do was produce light towards the red end of the spectrum - this is very useful as it can penetrate biological tissue more easily. The turning point came when Sergey Lukyanov at the Shemyakin Institute of Bioorganic Chemistry in Moscow, Russia, discovered a GFP-like protein in coral, which was termed DsRED.
’The red proteins from coral were larger and had a tendency to form tetramers, so they needed extensive modifications to make them user-friendly,’ Tsien says. This resulted in a new series of delicious sounding proteins with names like mPlum and mStrawberry - indicating the different shades.
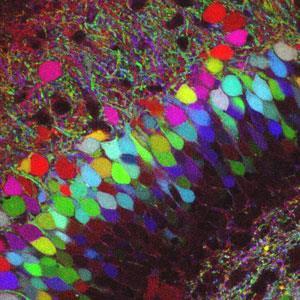
Perhaps the most vivid demonstration of the new family came when a Harvard University team, led by Jeff Lichtman in 2007, dyed neurons in mice in a kaleidoscope of 90 different colours - giving them the chance to look for unique patterns.
Bittersweet prize
The rules of the Nobel prize limit it to a maximum of three people. And, like many other important discoveries, more scientists contributed to the development of GFP. This year the man who cloned and sequenced the GFP gene, and who first imagined its potential, Douglas Prasher, missed out on this most prestigious of accolades. What’s much worse is that Prasher’s career was cut short. He failed to find funding for his research and dropped out of science entirely. He is now working as a bus driver.
’That Doug could not be recognised certainly lends a bittersweet element to the prize,’ Chalfie says. ’I think it must have been an agonising decision for the Nobel committee, and they could easily have given the prize to Doug instead of me.’
Tsien also acknowledges Prasher’s contribution saying: ’It’s a shame that Doug has not been recently in a position to do science. I hope all this publicity will result in Doug once again being able to do something for which he is really talented.’
Whether biology features too prominently in the Nobel prize for chemistry is a question raised almost every year. ’Our work certainly falls on the boundary line between chemistry and biology, but we didn’t do this work for the purpose of winning a Nobel prize - we did it because it was scientifically important and interesting at the time,’ says Tsien.
Chalfie agrees and places his work firmly on the border between the sciences. ’I think that the barriers between the different science departments are becoming more and more porous. And I think the Swedish Academy have certainly acknowledged that here.’ Without the chemistry of GFP, the vast range of important biological applications would have been impossible.
Further Reading
-
O Shimomura et al, J. Cell. Comp. Physiol ., 1962, 59, 223
-
D C Prasher et al, Gene , 1992, 111, 229
-
M Chalfie, et al, Science , 1994, 263, 802
-
R Heim et al, Proc. Natl. Acad. Sci. USA , 1994, 91, 12501







No comments yet