Harnessing the wisdom – and money – of crowds has rocketed in popularity in recent years. Clare Sansom looks at whether chemistry can join the gang
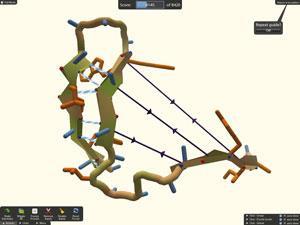
Structural biologist Mariusz Jaskólski of Adam Mickiewicz University, Poznan, Poland and his co-workers had a problem. They were trying to solve the crystal structure of a protease from the Mason-Pfizer monkey virus, which is closely related to HIV but is unique in that its protease crystallises as a monomer rather than the active dimer. Although they had obtained excellent x-ray diffraction data, none of the models available was close enough to be used as a template for molecular replacement. Success only came when the researchers challenged players of the computer game Foldit to generate models. Remarkably, it took a group of players only three weeks to produce several models that were good enough to solve the structure. ‘This unusual structure shows us that it may be possible to design molecules that prevent HIV protease from forming dimers, and that may be effective as drugs against Aids,’ says Jaskólski. ‘And we only managed to solve it with the help of these gamers.’
Foldit is the brainchild of David Baker of the University of Washington in Seattle, US, the main author of an acclaimed ab initio protein structure prediction program, Rosetta. ‘We realised that Rosetta needed more computer power than we had available and so developed a massively parallel version to run using “spare” computer time on volunteers’ home PCs,’ says Baker. Although most of the computers that run this variant, Rosetta@Home, are relatively low specification, there are now over 300,000 of them and they provide more power than the largest supercomputer.
‘Many of our volunteers became so interested in the science behind Rosetta that they wanted to be more involved, so we collaborated with computer games experts at Washington to develop a version that works as an interactive game,’ he explains. Foldit has scored notable successes besides the retroviral protease structure, including important contributions to enzyme re-design. ‘We have tens of thousands of players, but only a few of them will ever provide solutions that contribute to serious research,’ he says. ‘It is also interesting that few of the best players have “day jobs” as molecular biologists. I play it myself, but I’m not very good.’
Both Foldit and Rosetta@Home are examples of the phenomenon of crowdsourcing. This relatively new term, coined in 2006 by Jeff Howe of Wired magazine, is used to refer to projects that use the power of the internet to harness the resources of ‘crowds’ of people from outside the organisations where the projects initiate. James Surowiecki’s influential book, The Wisdom of Crowds, describes many occasions in which large, diverse groups of individuals have solved problems better than small numbers of experts, but these are only defined as true crowdsourcing if the web is involved.
Carl Esposti, chief executive of consultancy firm Massolution and founder of Crowdsourcing.org, divides the crowdsourcing ‘marketplace’ into a number of distinct segments. ‘There are micro-tasks, which are quick, simple and use only basic skills, expertise-based tasks and problem-solving or ideation tasks in which volunteers are polled for their ideas and opinions or for feedback,’ he explains. In this terminology, volunteer computing projects such as Rosetta@Home can be seen as a specialist type of micro-task with volunteers donating their computers’ time instead of their own. Crowdfunding projects are those that seek volunteers who are willing to donate their money rather than their time.
Phone home
The first volunteer computing project to attract large numbers of participants was SETI@Home, launched in May 1999. This project, which is still running, analyses radio waves from outer space for signals that suggest intelligent life; it has at times had over three million computers linked up. It also inspired the development of the first significant enterprise of this type in the life or chemical sciences. The Lifesaver screensaver was developed by Graham Richards and his co-workers at the department of chemistry at Oxford University, UK, in partnership with the US distributed computing company United Devices of Austin, Texas. Between 2002 and 2007, computers linked up to the screensaver ‘docked’ a billion molecules into the binding sites of 14 proteins that had been validated as cancer drug targets, generating thousands of potential leads.
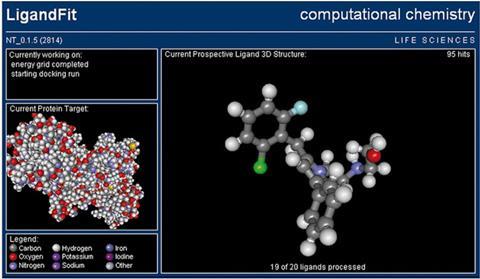
Later, Richards became a co-founder of the Oxford-based company InhibOx, which is exploiting both the technology and the hit compounds generated within this project. ‘The screensaver generated an enormous number of hits, many of which were “theoretical” molecules that could not be easily synthesised,’ says Paul Finn, chief executive of InhibOx. ‘None of these has yet produced a candidate drug that has entered clinical trials, but the screensaver project has nonetheless been invaluable: in particular, it has given us novel methods for dealing with enormous volumes of data.’
Potential volunteers now have many such computing projects to choose from. Most of these use a generic interface, the Berkeley open infrastructure for network computing (BOINC), developed by David Anderson of the University of California, Berkeley, US. Anderson is the author of the SETI@Home software, and BOINC is based on that original code. Users may link several projects to the BOINC interface simultaneously and decide how to partition their computers’ spare CPU cycles between each one.
Powerful backing
The computer giant IBM is a powerful supporter of the BOINC initiative via its World Community Grid (see Chemistry World, February 2011, p52). IBM works with scientists to adapt the software programs that govern their research projects, and then promotes the efforts widely, particularly within the company. ‘Since November 2004, over 600,000 users have donated over 650,000 CPU years of computer time to our projects, saving more than half a billion dollars,’ says Viktors Berstis, lead scientist at World Community Grid. Projects supported by the company include FightAIDS@Home, the Clean Energy Project (aiming to discover new materials for the next generation of solar cells) and the drug discovery project GO-FAM (Global Online Fight against Malaria). This is screening libraries of drug-like compounds against 18 potential drug targets from the proteome of the parasite Plasmodium falciparum, which causes malaria; details of one of these targets have been donated by InhibOx.

Once a volunteer has set up the BOINC software and linked it to one or more programs, they need have no further involvement in the scientific endeavour concerned. All the projects have websites, however, and some, including GO-FAM’s, include quite extensive educational resources to inform and inspire those who do want to know more. And it is a small step from that level of involvement to the ‘micro-task’ type of crowdsourcing. One type of human task that is easily adapted to large numbers of unskilled volunteers with relatively low involvement – and which is very difficult for computers – is that of pattern matching. This is the principle behind simple online games such as Galaxy Zoo, in which users classify images of galaxies based on their shapes, and now Cancer Research UK’s new Cell Slider project.
Diagnosing cancer often involves identifying potentially cancerous cells in images of large cell samples taken from tissue biopsies. ‘You don’t need to be a trained pathologist, or even know anything about cancer, to learn to distinguish tumours from normal cells,’ says Iain Foulkes, executive director of strategy and research funding at Cancer Research UK. ‘We developed the Cell Slider game with the help of Zooniverse, the company behind Galaxy Zoo, so volunteers could help us analyse our huge backlog of biopsies.’ Users are first taken through a tutorial to learn to distinguish the cells required, and regular users are monitored so more weight can be given to results from the most reliable players. And the game is proving popular, with the site attracting over 20,000 unique hits in the month following its launch on 24 October 2012.
Other organisations, including some large companies, have turned to crowds to solve more complex problems or provide expertise. Pharma giant Eli Lilly is a leader in this area and has spun out several ‘open innovation’ companies in the last decade, including InnoCentive, based at Waltham in Massachusetts, US. This links companies seeking solutions to problems with potential solvers via a website. The companies pay to register their details but may remain anonymous; solvers pay nothing, must formally register to receive details of each challenge, and receive cash rewards if they are successful. One of the most successful challenges was in synthetic organic chemistry: an anonymous solver came up with a novel, cheap and efficient synthetic route to a potential cardiovascular drug, and the drug is now in clinical trials.
Passing round the hat
Another site, Marblar, aims to turn the Innocentive model ‘on its head’ by providing scientists with a free space on the web to present solutions in search of problems (see Chemistry World, October 2012 p18). ‘Users come up with suggested uses for technical innovations and then vote on each one,’ says Richards, who is on the Marblar board.
Ideas and applications that have been linked and ‘liked’ by Marblar users may have something of a head start in attracting funding, particularly from less conventional funders such as ‘business angels’. It is a small step from there to inviting funding from the crowd directly. Since the 2008 financial crisis, funding from some traditional sources, both public and private, has been in short supply. Initiatives that seek to raise funds from large numbers of initially unconnected individuals are classed as ‘crowdfunding’. A recent survey of this emerging market segment by Crowdsourcing.org found that over $1.5 billion (£938 million) had been raised in this way in 2011.
Chemistry has not yet set its data free for the community
Esposti and his colleagues divided crowdfunding platforms into four types. In donation- and reward-based models, funders either contribute philanthropically or contribute to a cause in return for rewards or perks. Debt- and equity-based models are approaches where funders expect to earn interest and capital repayment or receive equity in the company or venture concerned. Crowdfunding attracts many small donations, so developers need to cast the net as widely as possible. ‘Linking with potential contributors through social media including Twitter and Facebook has become an essential component of any crowdfunding strategy,’ says Esposti.
Although Cancer Research UK is one of the best-funded medical research charities in Europe, it also has turned to crowdfunding, with the additional aim of linking its donors and its scientists more closely together. The strapline of its My Projects website is ‘Choose the cancer you want to beat’. This provides links to about 30 different CRUK projects – a small fraction of the charity’s total research portfolio – for potential donors to choose between. Despite that strapline, not all the chosen projects focus on a particular type of cancer. Donors are provided with information about their chosen projects that can be used for promotion in social media, and some choose to become much more closely involved. ‘A few of our researchers have developed very close links with individual donors, even inviting them to visit their labs,’ says Foulkes. ‘For this to happen, researchers need to be at ease with the media and able to describe their research in simple terms for non-experts.’
Opening up

It is rare for researchers, even in academia, to open their research up to the wider world to seek both criticism and collaboration. But Matthew Todd, a lecturer in organic chemistry at the University of Sydney, Australia, is one of the few who adopts this approach. He calls this open or citizen science, rather than crowdsourcing as it is strictly defined, although there is a clear overlap between the two. His entire group’s work, including the raw data in their lab notebooks, is posted on the web. Not only may anyone comment on this work while it is still ongoing, but anyone may contribute to the research, for example by synthesising molecules for testing.
Todd’s research, much of which is funded through standard Australian government grants, focuses on drug discovery for tropical diseases including malaria, so altruism is a clear motivation for participants. ‘I have been surprised to find that the majority of those making substantial contributions to these citizen science projects have been chemists from industry,’ says Todd. He protects his research through Creative Commons licensing rather than patents and dreams that this might become a norm in drug discovery, particularly for neglected diseases. There is certainly a historical precedent for this. ‘Both penicillin and the polio vaccine were discovered and developed without any IP protection,’ he says.
Take any comprehensive survey of crowdsourcing or crowdfunding – such as those produced by Crowdsourcing.org – and you will find fewer projects listed in chemistry than in many other disciplines. If crowdsourcing projects seek to ‘make communities of active stakeholders in research that affects them’, as Esposti suggests, it may be that chemistry projects are less immediately appealing to outsiders than those concerned with, for example, medicine, the environment or astronomy. ‘Chemistry projects, too, have huge potential, but chemists need to explain their projects and the value of contributions in terms that everyone can understand,’ he adds.
Todd, however, disagrees. Open data is a pre-requisite for open science, and too much chemical data is still locked behind paywalls in password-protected databases. ‘The agreement to keep human genome data free to all users has had incalculable benefits,’ he says. ‘Chemistry has not yet had its “human genome moment”, to set its data free for the whole community.’
Although chemists can be reluctant to open their lab notebooks to the world to take full advantage of the wisdom it has to offer, they seem to be more inclined to help those who are doing so. And with funding for research harder to secure, it would be less surprising if they embraced the challenges – and potential money – that crowdsourcing and crowdfunding has to offer.
Clare Sansom is a science writer based in Cambridge and London, UK.












No comments yet