For billions of years DNA has been life’s data storage medium. Now, scientists have used DNA to code and store their media and information, from all of Shakespeare’s sonnets to an audio recording of Martin Luther King’s ‘I have a dream’ speech.1
‘If you stick a CD in a box ‘round the back of the sofa it’s good for a couple of years but it might not work three years later,’ explains Nick Goldman of the European Bioinformatics Institute, just outside Cambridge in the UK, whose team performed the work. ‘In 10 years’ time, when you take the box out of the garage, you might not even have a CD player anymore.’
By contrast, DNA is long lived. Dried DNA can last for years, as Goldman explains: ‘They go drilling cores in Antarctica and the get bacteria out from 100m down in the ice. They reckon those are millions of years old and they get DNA out of them.’ That stability, along with the increasing developments in DNA production and sequencing make the idea of DNA information storage an attractive option.
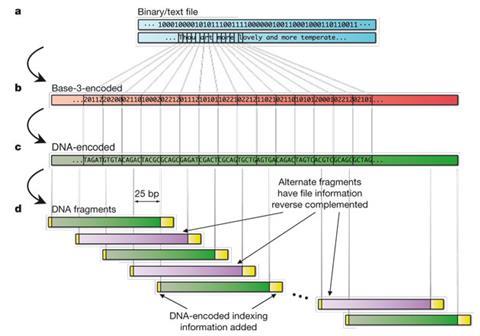
Even though we talk about the letters of DNA, there are only four bases (A, C, G and T), so there are not enough letters to spell out even a simple sentence. Instead, Goldman’s group converted data into binary and then each byte was converted using various codes and information theory algorithms into a DNA sequence that was very information dense but avoided runs of the same base, known as homopolymers. ‘Both the writing and reading techniques are more prone to making mistakes if there’s a run of the same DNA base in a row,’ explains Goldman. ‘If you’ve got two or three or four As – adenines – in a row it’s harder to read that back exactly right.
A technique whose time has come
In August of last year, a US group of scientists also published a paper demonstrating information storage in DNA.2 But while the two groups, working simultaneously, had some similar ideas their approaches are different. Goldman’s work focuses on reducing errors by using what are called error correcting codes, to avoid issues like homopolymers, while the alternative approach was careful to avoid sequences within the DNA that could result in secondary structures and folding within the polymer. It was these secondary structures that caused the only read errors in Goldman’s work and he is confident that further work can avoid these in future.
‘I think it’s very important and somewhat undervalued how important it is, especially for a new field, to have two groups that are independently doing it,’ says George Church, a geneticist at Harvard University, who is an author on the US paper. This new field, he says, is only really beginning now that the technologies are good enough to read the DNA.
But is DNA storage just for large archives? As the costs of DNA reading and writing technologies comes down both Church and Goldman suggest that DNA might be the read only medium of choice. In fact, both Goldman and Church suggest that DNA-based information storage might just be the commercial push needed to really lower the cost of DNA writing, much as medicine has pushed down the cost of sequencing.
For Sriram Kosuri, who worked with Church on the 2012 paper, the value of both of these pieces of work is that they get people to think differently about data storage. ‘We’re reaching a limit on how much we can store on a surface,’ he explains, mentioning the current world record of a 12 atom transistor. ‘Both of these papers blow that out of the water in terms of density and longevity.’ And that polymer might not be DNA, but another polymer that’s easier to read and synthesise and less prone to enzymatic digestion. ‘We’re using DNA in its rawest form, but I think in the future we can imagine using something very different,’ he concludes.






No comments yet