New technique uses DNA origami to provide visual read out of the most common form of genetic variations
DNA origami and atomic force microscopy have been combined to reliably detect and image single nucleotide polymorphisms (SNPs) - the most common form of genetic variation in the human genome. The advance could contribute to the growing field of personalised medicine, and illustrates the potential of DNA origami for diagnostic applications.
SNPs are variations in a single nucleotide (A, T, C or G) in a stretch of DNA, and can determine how individuals might be susceptible to certain diseases and respond to pathogens, chemicals and drugs. ’In terms of genetic variations and potential medical applications, you want to know that certain things are present in genomes and this is a way to get at them,’ says Ned Seeman who led the study at New York University, US.
Detecting SNPs usually requires expensive DNA microarray techniques that label sections of DNA where SNPs occur. However, Seeman’s approach differs in that there is no need for labels, using a technique based on a visual read out.
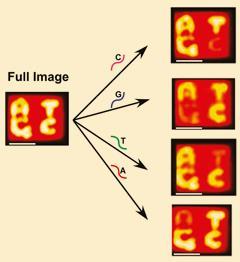
First the team created the letters A, T, C and G on a DNA origami tile 100nm square. They did this by using single staple strands of removable DNA that, when exposed to UV light, became two different strands - one holding the DNA origami together while the other revealed tiny bumps that acted like pixels to show a pattern of an individual letter.
The bumps making up each letter, however, also have distinct binding sites that are complementary to the nucleotide they represent. So for example, the strand that displays a T shape has sites that are complementary to thymine. This means that when a DNA strand containing an SNP involving thymine is inserted into the system, it binds only to the strands that make up the T pattern, thereby removing it while the other letters remain visible under AFM.
Paul Rothemund at California Institute of Technology in Pasadena, US who pioneered DNA origami in 2006 thinks the paper is ’really great’ because it showcases the idea of DNA origami as a ’DNA nanochip’. ’Seeman’s paper is the first example of a practical application for which DNA origami can really be used as a custom instrument for answering biological questions,’ he comments. ’It encourages people to quit messing around and to really develop complex biological applications.’
But Rothemund is unsure if a new SNP detection method is really necessary as full genome sequencing could become cheaper and more widely available for individuals, possibly within a decade. ’With a full genome comes all the single nucleotide polymorphisms, but more importantly everything else you care about for genetic medicine.’
Until that happens, however, Seeman reckons that his new technique could offer an alternative, and possibly more reliable, SNP detecting solution. ’We’re putting this technique out there and it’s up to the community to decide if they think it’s better than other methods,’ he says. ’What we’re really waiting for is someone to develop a miniaturised AFM and then you will be able to just take our technique into the field and all you would need to know is how to read A, T, G and C.’
James Urquhart
Interesting? Spread the word using the ’tools’ menu on the left.References
H K K Subramanian et al, Nano Letters, 2011, DOI: 10.1021/nl104555t





No comments yet