The Venter Institute unveils its man-made microbe - a chemically synthesised genome housed in a naturally grown cell
A chemically synthesised chromosome has for the first time been transplanted into a cell to produce a synthetic bacterium. The advance provides a basis for making organisms designed from scratch and represents a major step towards applications in biofuels and chemical synthesis through synthetic biology.
The man-made microbe is the work of a team led by Dan Gibson and genome sequencing pioneer Craig Venter, at Venter’s institutes in Rockville, Maryland and San Diego, California, US. Besides a few genetic ’watermarks’ encoded by the team, its genome largely duplicates that of a goat parasite called Mycoplasma mycoides. Grown in a dish, the synthetic version looks much like the original and, like its natural counterparts, is capable of self-replicating.
’For the past 15 years, the genomes of many organisms have been sequenced and deposited in databases. We call this digitising biology,’ says Gibson. ’We now show that it is possible to reverse this and synthesise cells starting from this digitised information.’ The aim, essentially, is to be able to carry all the information required for making life around on a memory stick.
Actually making it, however, is rather trickier. The M. mycoides genome contains more than a million ’letters’ of code - nucleotide base pairs (bp) - but current DNA synthesising technology can only string together a few thousand at once. Thus, Gibson’s team exploit the ability of yeast to stitch together small DNA fragments using enzymes. Since announcing the first synthetic genome in 2008,1 they have been attempting to create synthetic life by transferring bacterial genomes assembled in yeast into naturally grown host cells.
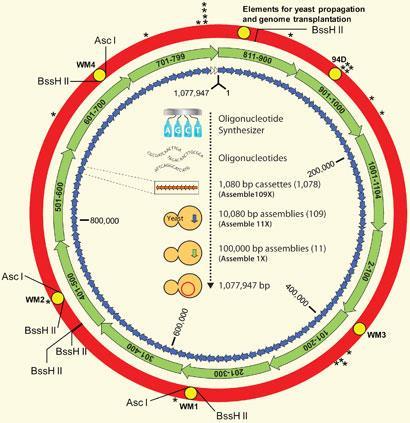
In the present study,2 over a thousand short (1,000 bp) pieces of DNA were chemically synthesised and joined by overlapping ends - as Gibson explains, the yeast recognises the overlap in the code and sews them together. Larger segments of first around ten and then a hundred thousand base pairs were combined to produce the full length synthetic genome, which was finally transplanted into the host, another bacterium of the same genus called M. capricolum.
So Gibson’s work provides the tools for creating artificial life, but what about the blueprints? Designing an organism from scratch requires an understanding of what each gene codes for. But we’re getting there, according to Paul Freemont of the Centre for Synthetic Biology and Innovation at Imperial College London, UK.
’The extrapolation of course is that you could actually design genomes and there’s a lot of ground to cover before then, but we do understand quite a lot about what all the genes code for, so I think it’s a very major step,’ says Freemont. ’More likely would be to try to make a minimal synthetic genome, containing all the basic properties of a living organism, which would allow you to put other types of gene circuits into it, like biofuels or fine chemicals.’






No comments yet