Philip Ball explains how creative chemists are teaching molecules some new tricks
In a highly prescient book,1Jean-Marie Lehn remarked that the increasing complexity of synthetic chemical systems should one day extend to ‘chemical “learning” systems that … can be trained, that possess self-modification ability and adaptability in response to external stimuli’. If you can learn, you can evolve – and in this way the complexity of artificial chemical systems might recapitulate the development of life on Earth, which presumably progressed from simple copying mechanisms to adaptive assimilation and transmission of information.

Computer science and neurobiology have sharpened the notion of what learning is and how it happens, making it seem well within the reach of the most rudimentary information-processing networks. The classic view, due to psychologist Donald Hebb, invokes reinforcement of communication pathways between the nodes of a network – neurons connected by synapses, say – so that particular stimuli reproducibly activate particular collectives of nodes. This ‘Hebbian’ model is sometimes crudely expressed as ‘cells that fire together, wire together’.
That view has helped to develop neural theories of learning and to embody them in artificial neural networks in silico. But a freely diffusing chemical system doesn’t have a well-defined spatial topology connecting the interacting molecules. However, the image of molecular systems as networks has become increasingly familiar in cell biology. The specificity of enzyme reactions, for example, makes it possible to regard biochemical processes as networks of reagent nodes linked to nodes that co-react. The same is true in genetics: forging a connection between genomes and phenotypes is now largely seen as a matter of mapping out the network of gene interactions mediated by regulatory proteins and RNAs.
The prevailing picture is stimulus and response: the interaction pathways might be complex and nonlinear, but the idea is that a particular input – a viral invader, say – stimulates a predictable outcome. Even here, however, learning may be an option: genetic networks can be conditioned in ways that don’t demand changes at the genotypic level. Just think how the immune system ‘knows’ how to respond to challenges encountered years earlier.
Yet learning can be designed more explicitly into gene networks. In 2008, a team in England and Germany outlined a theoretical design for a gene regulatory network that could be written into plasmids and inserted into Escherichia coli, enabling these organisms to display associative Hebbian learning.2 The network incorporates feedbacks that allow it to be trained, so that a particular (but arbitrary) chemical stimulus elicits a learnt response such as producing certain proteins.
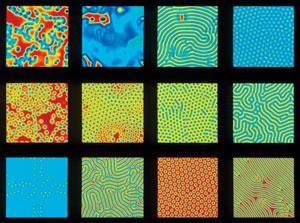
Peter Banda and Christof Teuscher of Portland State University in the US believe that learning can occur in even simpler systems. Last year they outlined a theoretical chemical system, the ‘perceptron’: a simple learning unit that can be trained to integrate a given combination of input signals into a particular output.3 The reacting species can act as catalysts or as rate-lowering inhibitors, and learning depends on feedbacks that tune the concentrations of these catalyst–inhibitors, driving the input-to-output reactions. The idea is related to the activator–inhibitor systems postulated by Alan Turing, which are well known to give rise to complex behaviours such as pattern formation.4 Indeed, chemical kinetics similar to Turing’s model have already enabled purely chemical systems to mimic intelligent behaviours.5
With an appropriate choice of rate constants, the chemical perceptron could, for example, learn to function as any of the standard logic gates in normal Boolean circuits. Now, in collaboration with US computer scientist Darko Stefanovic of the University of New Mexico, the team has simplified the model so that it becomes possible to imagine implementing it in a real chemical system.6 In short, the researchers encode binary signals in terms of a threshold concentration of a single species, rather than two distinct species, halving the ingredients.
This scheme looks more practical, so it’s worth considering what chemical systems might be used for real ‘wet’ experiments. One possibility is enzymes; another is DNA systems in which single strands can exchange in double-stranded complexes. Eric Winfree at the California Institute of Technology has shown these reactions can be used to represent any chemical kinetic scheme.7
If it works, chemical learning would enable programmable reactions, trained for a particular output without having to be hardwired at the level of individual reagents. There would be few more dramatic examples of Lehn’s vision of chemistry as an information science.










No comments yet