Tissue samples inflated to four times their size to overcome microscopy limitations
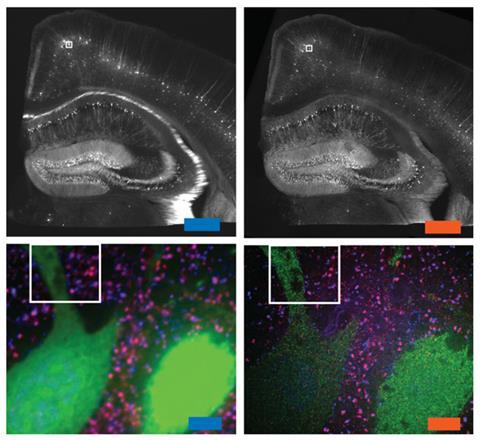
A new microscopic technique that magnifies specimens by blowing them up like a balloon could make it easier to produce high resolution images of cells and tissues. The team behind the technique showed it could be used to image brain tissue using an ordinary confocal light microscope at around four times the resolution that would normally be possible.
The technique – dubbed expansion microscopy – differs from existing super-resolution microscopy methods, including those pioneered by the winners of last year’s chemistry Nobel prize. Rather than having to create special microscopes or image samples many times, expansion microscopy works by making the sample physically bigger using the swellable polymer sodium polyacrylate – a super-absorbent material used in disposable nappies.
‘Our method lets you image large, intact, 3D structures with nanoscale precision, [which is] very helpful for mapping large brain circuits with nanoscale precision, imaging tumours with resolution precise enough to see nanoscale signalling cascades and other problems in biomedicine where you want to look at large tissues with great accuracy,’ says Ed Boyden from the Massachusetts Institute of Technology, US. ‘And it also works with the conventional microscopes that most groups already have.’
The idea was born out of frustration in Boyden’s group, who work on mapping neural circuits in the brain. ‘We had been half-jokingly kicking around the idea of making the brain bigger for quite a long time,’ Boyden says. After struggling to get detailed enough images with electron microscopy and super-resolution microscopy they decided to give the idea more serious thought. ‘Amazingly, the basic principle of tissue expansion started working right away,’ says Boyden.
What’s the big idea?
In expansion microscopy, the polymer is first embedded in the tissue by infusing its precursors (the monomers sodium acrylate and acrylamide, and cross-linker N,N’-methylenebisacrylamide) into the sample and triggering free radical polymerisation with an initiator. Then, a protease enzyme is used to homogenise the tissue–polymer composite and water is added, making the polymer swell to over four times its original size. ‘Because the polymer chains are winding their way through the tissue, as they grow, they take the tissue with them. Thus, the tissue sample itself becomes bigger,’ says Boyden.
The group also developed fluorescent labels that can be used to highlight specific cell components. The labels contain a methacryloyl group that is incorporated into the polymer, as well as a coloured fluorophore and an oligonucleotide capable of binding to a target sequence to highlight specific proteins.
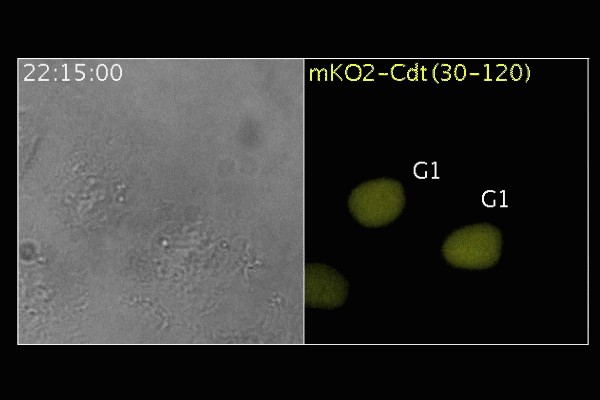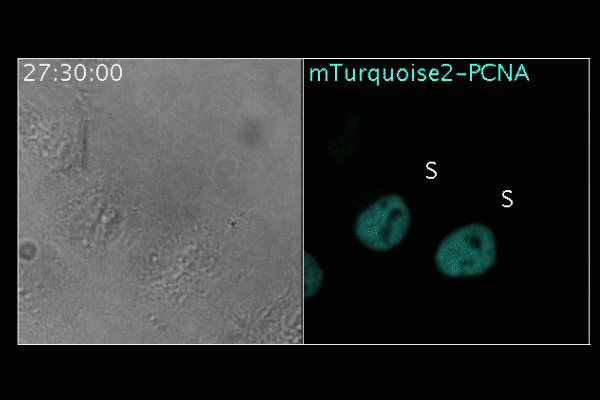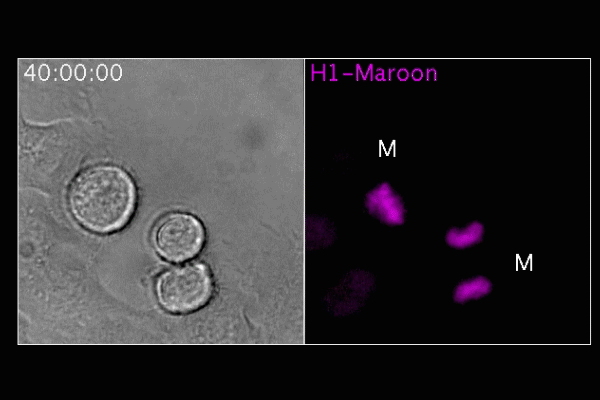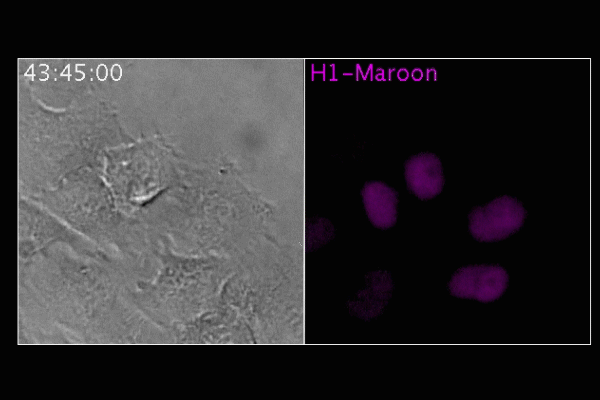
By blowing up samples, Boyden and his colleagues were able to image microtubules within cultured cells at high resolution using a standard confocal light microscope. When compared with super-resolution microscopy images of the same structures, the expansion microscopy images were just as detailed, and in some cases the resolution was even sharper. The team also imaged mouse brain tissue using the technique, and were able to distinguish structures at around 70nm resolution – far beyond the normal limit of 250nm for the microscope they were using. Synthesising different labels targeted to various proteins in the brain allowed them to produce multicolour images.
Harsh treatment
‘The images are pretty convincing and the quality seems comparable to the existing super-resolution microscopy, without having to purchase expensive equipment,’ says Violaine Sée, co-director of the centre for cell imaging at the University of Liverpool in the UK, ‘It is surprising that by expanding the sample they don’t get any distortion.’
She is sceptical, however, that the technique will be fully embraced by biologists, because of the harsh way in which the samples must be processed. ‘What they image in the end – microtubules, for example – is only a ‘ghost’ of the microtubules, because they were digested [by a protease] after the labelling,’ she says. But she adds that it may be useful for some approaches, such as light sheet microscopy, that already involve heavy processing, as people will be less reluctant to try new preparation methods.
Boyden and his colleagues have already set their sights on even higher resolution. ‘We are working to make the expansion ratio bigger, meaning that objects will become even bigger than before,’ he says. They are also working on tweaking the method so it can be combined with different assays for in situ analysis. ‘These will be challenging,’ Boyden says, ‘but may eventually allow imaging of many of the components of a biological system, with nanoscale precision, throughout the entire system.’









No comments yet