Many drugs that are normally used to treat non-infectious diseases like cancer, diabetes and depression can kill bacteria through modes of action that are different from those of standard antibiotics. The US-based researchers behind this finding say that it opens up new possibilities in the battle against antibiotic resistance.
‘Our findings uncover molecules that can be chemically modified and improved for the potential development of new antibiotics,’ says Amir Mitchell from University of Massachusetts Chan Medical School who led the study. He and his colleagues evaluated the toxicity of a large number of drugs on thousands of Escherichia coli strains mutated in different genes to determine the functional relationships between antibiotics and non-antibiotics.
The researchers identified 186 drugs with antibacterial activity from a pool of over a thousand different compounds. ‘Toxicity was tested in liquid growth media, which becomes cloudy once bacteria grow in it,’ explains Mitchell. ‘If the media remains clear, the drug is toxic.’ The team then used a screening method they developed in 2020 to examine the cellular pathways involved in this activity. In total the team mapped around two million gene–drug interactions.
‘We grew a collection of thousands of different bacterial mutants with each of the drugs that have antibacterial activity and used DNA sequencing to calculate the frequency of each mutation before and after the drug,’ says Mariana Noto Guillén, who did many of the experiments. ‘If a mutation gives resistance to the drug, that specific mutant strain will grow much more than all other mutants in the collection, while if the mutation causes sensitivity, the frequency of the mutant will be greatly reduced.’
The scientists carried out a network-based analysis – using their own computer code and algorithms – to unveil similarities among the studied drugs. ‘The computational analysis was designed to identify, for each drug, a list of mutants that are very sensitive or very resistant to it,’ mentions Mitchell. ‘If two individual drugs have very similar lists, they likely kill bacteria with a similar mechanism.’
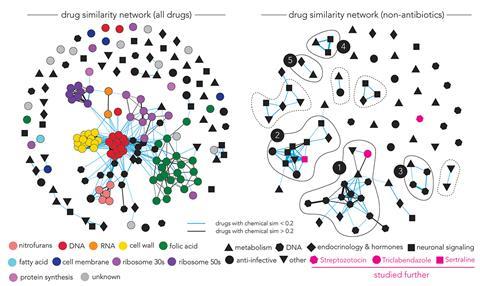
‘The researchers observed that functionally similar antibiotics had similar drug–gene interactions, as one would expect,’ comments Jon Stokes at McMaster University in Canada, who wasn’t involved in the study. Indeed, they found that all antibiotics that target the cell wall were clustered together and well-separated from other types of antibiotics such as those interfering with protein translation.
But they also found that non-antibiotic drugs with antibacterial activity didn’t form clusters with known antibiotics, confirming the different modes of action of these compounds. About half of the non-antibiotics formed clusters with each other, revealing possible shared and unexplored mechanisms. Understanding these mechanisms could guide the design of new antimicrobials, points out Mitchell.
The team finally analysed the effect of the drugs on the efflux pumps of E. coli. Efflux pumps are transport proteins that free bacteria from toxic molecules. ‘Genetic perturbation of the efflux systems resulted in sensitivity to both antibiotics and non-antibiotics alike,’ says Stokes. ‘This suggests that non-antibiotic drugs that display antibacterial activity may promote mutations in efflux systems, which could confer resistance against conventional antibiotics.’ Mitchell notes that these observations were made in vitro and so careful in vivo studies are still required to confirm this.
References
1. M N Guillen et al, Science, 2024, DOI: 10.1126/science.adk7368







No comments yet