UK scientists have stumbled across a molecule that ties itself up in knots, echoing the folding of proteins
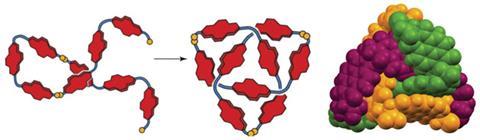
If knotting up molecules is so difficult why not take a tip from nature and get the molecules to do the work for you? This, it turns out, is not such an outlandish idea after Cambridge University researchers found one of their molecules was indeed tying itself in knots.
Jeremy Sanders’ group weren’t looking for knots, they were continuing their work on dynamic combinatorial chemistry, setting up a reaction mixture with components that undergo reversible chemistry and leaving them to equilibrate to the most stable product. As Sanders explains, ‘the whole idea is to set reactions up and discover something unexpected’. And lo and behold they did.
In this experiment the group used naphthalene diimide (NDI), linked together with hydrophilic alanines and capped with cysteine residues. This results in molecules that have both hydrophobic and hydrophilic sections, and the cysteine residues can make reversible disulfide links. ‘To anthropomorphise for a moment,’ Sanders says, ‘the building blocks find themselves in water and they’re not very happy, so they look around to see what they can bind.’ But all they have to bind to is another hydrophobic NDI. As disulfide linkages are formed, making longer chains, the NDIs try to bury themselves away, while the linkers try and stay in the water. Disulfide links are broken and formed while the chain tries to twist into the most stable form. Mostly this means that simple cyclic monomers and dimers are formed. However, during separation Sanders’ team noticed a tiny third peak, indicating the presence of a third species.
Amplifying the hydrophobic effect by using a higher concentration of the building block, or adding salt, increased the size of that peak. Mass spectrometry showed the compound responsible was the right size to be a trimer, but it was eluted too quickly for it to be a simple macrocycle. Perhaps it was a knot.
Knotty chemistry
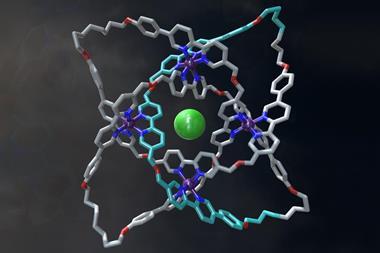
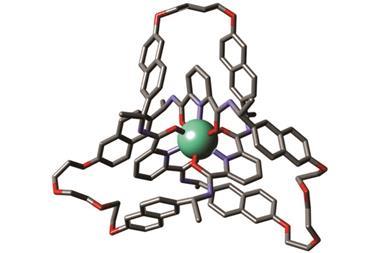
Further investigation confirmed that the molecule had indeed made a trefoil knot – a knot with three loops that cannot be untied without cutting it open. Back in 2010, chemists first managed to make a molecular trefoil knot, but that required a metal ion to act as the template. Over in the biological world, proteins often knot themselves up and, as Sanders explains, his team seems to have accidentally used the same method that nature does. Their evidence suggests that the NDI building blocks make a trimer and that these then knot themselves up. Once in the knot, the molecule is in a thermodynamic well, and it stays there.
‘The Sanders’ knot is fascinating,’ says David Leigh from the University of Edinburgh in the UK, who made the first molecular pentafoil knot last year. ‘One wouldn’t normally expect the most stable form of a closed loop to be a tight knot … but for this molecule the hydrophobic effect provides a powerful driving force.’
Leigh adds that he thinks this discovery has once again shown dynamic combinatorial chemistry to ‘be a powerful experimental tool for generating unexpectedly complex structures’. Sanders agrees, adding: ‘Once you’ve discovered something unexpected, and this is very beautiful and very astonishing, it suddenly opens a door to a whole new world.’ Currently, he’s keeping his cards close to his chest about what’s behind that door. ‘What I can say,’ he hints, ‘is this is the first knot we’ve made, but it won’t be the last.’






No comments yet