A new RNA editing tool is opening up new ways to develop novel RNA therapies, speed up engineering of human viruses and test drugs. In a proof of concept experiment, researchers used a cutting enzyme – an endonuclease – and a programmable RNA repair enzyme to modify an RNA virus genome.
Crispr is famed for its precision and efficiency in editing DNA. It’s a dual system that uses repeat sequences of guide RNA (Crispr) to lock onto a target piece of DNA, which is then cut by a Crispr-associated (Cas) protein. It is a strategy that archaea and bacteria deploy to recognise and slice up foreign DNA from viral enemies.
But Crispr has proved challenging to adapt to RNA. Editing an RNA virus requires transcription into DNA, changes are then made using readily available DNA engineering tools and then the new sequence is transcribed back into RNA.
Now, researchers at Montana State University, Bozeman, have tapped a type III Crispr system in a dairy industry microbe, Streptococcus thermophilus. Just like in DNA editing, the RNA is cut by a Crispr-guided nuclease guided to the target sequence by RNA. Once cut, synthetic splints of DNA bring the two RNA strands together for repair by a viral ligase enzyme.
‘There are around 40 Crispr subtypes, but only a few have been used in the same way as Cas9,’ says Blake Wiedenheft at Montana State. ‘Ironically, the type III Crisprs are amongst the most abundant in nature but haven’t been repurposed much in biotech.’
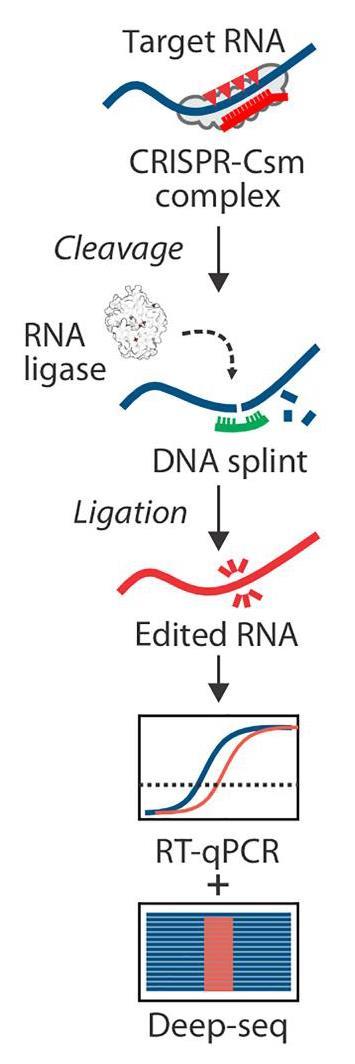
The group made programmed deletions in the genome of a Sindbis virus. This virus had a green fluorescent reporter gene engineered into its RNA, and after editing that targeted the reporter the virus stopped fluorescing demonstrating that the Crispr system was working properly.
The new advance could make it easier for scientists to perform loss- or gain-of-function research in RNA viruses, changing viruses to see how their virulence is affected, for instance. Or to knock out part of a viral genome to determine how a virus might circumvent a new antiviral drug.
Wiedenheft’s group’s work was inspired by an unusual variant of Sars-CoV-2, actually identified in Bozeman, that was missing the ORF7a gene. They wanted to confirm that this deletion caused its defects in replication by excising ORF7a from a Wuhan strain, but they struggled to carry this out leading them to search for a new way to edit RNA. ‘The most valuable and immediate use of this technology will be to evaluate different functional aspects of viral genomes,’ says Wiedenheft. However, the new system still needs to be more efficient and precise before it will find more widespread use.
The system could have many other applications. ‘You could directly modify an RNA virus,’ says Volker Thiel, a virologist at the University of Bern in Switzerland, to discover how easy or difficult it is for that virus to escape a drug. ‘You target a region where you think resistance might emerge to a new drug and then find out using imprecise changes which mutations confer resistance to it.’ A map of resistance mutations would be helpful in drug development or when choosing treatments for patients.
There is potential for creating new treatments too. ‘The therapeutic applications could include editing and fixing single-based DNA disorders without having to modify the genome,’ notes Darrell Green, a molecular biologist at the University of East Anglia, UK, with an interest in RNA silencing therapies. ‘Once the RNA is gone, there is no real trace like there would be in DNA editing.’
Some bone tumours produce a mutant protein. ‘RNA Crispr editing could restore the wild type by changing the mutant back – avoiding having to replace the wild type in its entirety,’ Green says.
Some people have described Crispr as the democratisation of genome engineering,’ Wiedenheft says. ‘This RNA editing approach now allows us to do the same thing at the RNA level. This is just the beginning.’
References
A Nemudryi et al, Sci. Adv., 2023, DOI: 10.1126/sciadv.adj8277















No comments yet