The processes underpinning how solvent and solutes molecules interact are fundamental, but still mysterious. Philip Ball investigates
The performance of an electrochemical capacitor; stereoselective catalysis for drug synthesis; how protein chains fold up. Beyond the fact that these are all chemical problems, they seem to have nothing in common with each other.

Yet they do. They are all concerned in one way or another with the question of how solvent molecules interact with solutes and surfaces. And in each case, mastering the problem is currently hampered by an inadequate understanding of this issue of solvation. Because solvation has never been acknowledged as a distinct branch of the molecular sciences, there’s no general picture of what it entails.
An international initiative called Resolv, launched in June and based at the University of Bochum in the Ruhr region of Germany, hopes to change that. It aims to establish solvation science as a discipline in its own right, with principles that should apply to chemical processes in areas as diverse as corrosion, molecular biology and green chemistry.
‘I believe that the importance of solvation science is similar to that of neuroscience or materials science, but that this has not yet been recognised,’ says theoretical chemist Dominik Marx, who is coordinating the solvation and interfaces program of Resolv, one of its three key focuses. ‘I think that we can find common viewpoints that link hitherto unconnected phenomena.’
Resolv has big ambitions. It is one of 12 ‘clusters of excellence’ picked by the German research council from 10 times as many applicants, and will receive a total funding of €28 million (£24 million) over a five year period, along with a lump sum of €44?million from the German Council of Science and Humanities for constructing a new experimental research facility at Bochum. It includes several regional partners, and has set up worldwide collaborations with institutions ranging from the University of California at Berkeley in the US to the Weizmann Institute in Israel and the Indian Institutes of Technology at Bangalore and Kanpur.
‘Solvation science as such hasn’t previously been identified as a distinct field at the interface between chemistry, physics and biology,’ says Martin Weik, a biophysicist at the Institute of Structural Biology in Grenoble, France. ‘It is perhaps this complex interdisciplinarity that prevented the establishment of a well-defined field so far.’ He calls Resolv ‘unique, timely and exciting’, saying that it ‘has great promise to become a global network and platform for understanding solvation’.
Victim of success
How substances behave in solvents is an ancient problem. Interest in solvation goes back at least to the alchemical idea of a universal solvent (or ‘alkahest’) that could dissolve anything. Its attraction was that, by dissolving everything into its elemental constituents – or so it was believed – it could potentially turn any substance into any other. Solvation was thus the key to transformation.
Solvation underpinned the origins of strongly applied fields such as metallurgy, colloid science and organic chemistry. Some of the basic rules governing the thermodynamics of solutions were established by early physical chemists in the 19th century, while in the 1920s Peter Debye and Erich Hückel in Zurich worked out a theory of how ions are dispersed in water. ‘Solvation science was real big around the turn of the century, thanks to people like Arrhenius, van’t Hoff, Ostwald and Debye’, says physical chemist Pavel Jungwirth of the Institute of Organic Chemistry and Biochemistry in Prague in the Czech Republic. ‘Then the field fell out of fashion – partially being victim of its own success, with the main issues considered solved and a few remaining things swept under the rug.’
Many textbooks only make passing references to solvents
Yet that understanding was in fact highly limited, mainly to ions in dilute solution. ‘The general theories of close-to-ideal systems are reasonably well established, but the devil is in the detail,’ says Jungwirth. ‘For example, physiologically relevant solutions are far from ideal.’
As a result, many of the solvation issues that chemists now face routinely, such as the effects of solvent on reaction rates, still lack a comprehensive theoretical framework. According to the authors of a recent textbook on solvents in synthesis, ‘many inorganic and organic chemistry texts only make passing references to solvents, or worse still, fail to mention that a given reaction takes place in a particular solvent at all’. The choice of solvent in organic chemistry relies on some rules of thumb, often based on sound but qualitative principles, coupled to a certain degree of trial and error. And of course there is no unique way to define ‘rightness’ anyway, because there are so many factors to consider: cost, volatility, toxicity, thermal and chemical stability and so forth.
Why should the time now be ripe for solvation science to be declared a subject in its own right? ‘If you want to unify something that has to do with molecules, you must understand things at the molecular/atomistic level, which allows for a bottom-up strategy,’ says Marx. ‘And when it comes to the molecular viewpoint, we have seen dramatic advances in both experiment and simulation in the last decade or so.’

Martina Havenith, Resolv’s principal coordinator, agrees. ‘Simulation and experiment have come closer to each other, and we are now able to approach experimentally solvent structure and solvent dynamics on a molecular level,’ she says. New techniques have been developed to address these issues, such as sum-frequency generation spectroscopy, terahertz spectroscopy and synchroton-based diffraction methods. Such findings can now be compared with the predictions of computer simulations that include many solvent molecules modeled explicitly, rather than as some uniform background medium. ‘We now have technology, primarily spectroscopic and imaging techniques and molecular simulations, which are putting solvation science on a firm molecular basis,’ agrees Jungwirth.
Life’s solvent
The molecular granularity of solvents seems crucial to understanding many questions of solvation. Nowhere is that more true than in the hydration of biomolecules. In 1959 the biochemist Walter Kauzmann proposed that the hydrophobic force – the attractive interaction between water-repelling solutes in water, such as the hydrocarbon side-groups of some amino-acids – can be explained on the basis of how water molecules are arranged around them. Since the hydrophobic force is one of the main drivers of protein folding and interactions between proteins, not to mention a central factor in the formation of lipid membranes and in the binding of many natural substrates and drugs by their receptor proteins, it is of central importance in molecular biology. Kauzmann drew on the earlier suggestion that water molecules hydrating hydrophobic particles will arrange themselves into a more tightly bound and orderly arrangement than in the bulk liquid – a kind of ‘iceberg’ – in order to preserve the hydrogen bonds between them. When two hydrophobic particles come together, said Kauzmann, some of this ‘structured water’ in their hydration shells is released into the bulk as the hydration shells overlap. That liberation of structured water supplies an entropic gain that makes the aggregation of particles favourable in free-energy terms.
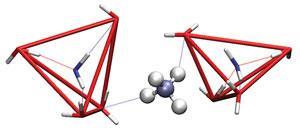
It is now widely recognised that Kauzmann’s iceberg model of hydrophobic hydration is too simplistic: water molecules in the hydration shell are not necessarily more ordered, let alone frozen into ice-like immobility, although some studies suggest that they might rotate somewhat more slowly than in the bulk.1,2 So it’s still not clear exactly what causes the hydrophobic force. In fact, chemist David Chandler, of the University of California at Berkeley, and his coworkers believe that there may be more than one mode of hydrophobic hydration, depending on the size of the solute particles. Above about 1nm, he says, water molecules can no longer just accommodate the interloper by rearranging themselves to form a cavity while preserving hydrogen bonding, but must accept the inevitable loss of some hydrogen bonds at the interface with the hydrophobic surface.3
This loss of hydrogen bonding at extended hydrophobic surfaces can lead to a situation in which it becomes thermodynamically favourable for all the water between two such surfaces to clear out in a concerted flight as the surfaces approach, leading to an abrupt ‘drying transition’ that snaps the surface together.3 There is some evidence that this can happen in some cases of protein folding and aggregation, but it is not clear how general the phenomenon is.4,5 One thing is clear, however: the details of what goes on cannot be understood or predicted on the basis of any simple and universal principles, but depend on the precise chemical composition and shape of the surfaces. This means that hydration needs to be modeled using explicit, individual water molecules – which is computationally challenging for a large solute molecule like a protein.
It is not easy to probe such complex effects experimentally either. ‘We need experimental techniques which cover the different timescales of solvation dynamics,’ says Havenith. But she says that a battery of methods for doing that is now available. ‘Scattering techniques, such as neutrons, give access to the hydration structure. NMR and dielectric spectroscopy probe the slower part of the hydration dynamics in the nanosecond and microsecond range,’ she says. ‘New laser techniques, such as femtosecond fluorescence, two-dimensional infrared terahertz and optical Kerr spectroscopies, now yield information on faster time scales.’
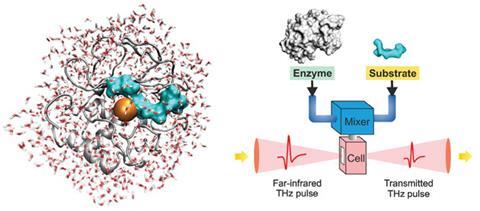
Havenith has been developing the relatively new method of terahertz spectroscopy, which is able to probe the collective motions of the water network on timescales of less than a picosecond (the typical lifetime of a hydrogen bond in bulk water). Her studies have shown just how amazingly complex and subtle biomolecular hydration can be. In collaboration with Irit Sagi at the Weizmann Institute in Israel, she looked at changes in the dynamics of hydration water as a zinc metalloprotease enzyme (involved in restructuring connective tissue during tumour genesis) binds its substrate. They found that as the substrate approaches the binding site, the water in the intervening space seems to become more sluggish, in rough proportion to its proximity to the binding site.6 This slower water dynamics in turn slows down the vibrations of the substrate so as to help guide it into place. In this way the enzyme seems able to ‘tune’ the character of the hydration water to help it do its job.
With that in mind, Weik hopes that ‘drugs can be rationally designed that act by modifying water and the solvent properties of their targets’. Many proteins do exploit individual water molecules for binding their substrates, using them as hydrogen-bonding bridges. More generally, it has become clear that hydration water plays a key role in keeping proteins and other biomolecules ‘active’, holding them in shape while injecting fluctuations that keep them supple enough to bind and release their targets. The increasing focus on these dynamical aspects of biomolecular behaviour has motivated the inclusion of water in the picture, according to Weik. ‘I think the field of molecular biology is now ready to fully integrate solvation aspects into their molecular understanding,’ he says.
Ion power
Another of nature’s tricks is to arrange water molecules into hydrogen-bonded chains which act as ‘wires’ along which hydrogen ions can be conducted by flipping of the hydrogen bonds, shifting the ions quickly over long distances as required in many enzymatic reactions. Such proton transfer is just one example of the ways in which solvation science can bridge very different areas of chemistry. ‘Charge transfer is essential not only to many processes in biochemistry but also in liquid-phase catalysis and electrochemistry,’ says Marx. Indeed, ion solvation is central to the processes of corrosion, mineral weathering (and thus the biogeochemical cycling of elements), electrocatalysis and battery technology, desalination, and nerve signaling, to mention just a few examples.
‘Electrochemical reactions are the basis for energy conversion or storage processes such as fuel cells, electrolysers and batteries,’ says Wolfgang Schuhmann, an electrochemist at Bochum and a member of the Resolv team. ‘If we want to develop these electrochemical systems based on fundamental knowledge about reactions at electrified interfaces, understanding the impact of solvation processes is indispensable.’ For example, he says, ‘in corrosion a metal atom gets oxidised and has to be dissolved – a process that clearly involves solvation. Lithium-ion intercalation in batteries is formally the opposite of corrosion: a solvated lithium ion first has to strip off its solvation shell.’ Until now, says Schuhmann, these solvation and desolvation steps have not been well understood, largely because the interface of an electrode and electrolyte is so complicated.
Because solvation surrounds an ion with a cloud of loosely associated solvent molecules, it gives ions an effective size considerably bigger than their ionic radius. This has profound consequences for how ions behave in very small confining spaces such as nanopores. For example, some desalination technologies are exploring the use of carbon nanotubes. Below diameters of about 8Å, water can pass through the tubes in hydrogen-bonded single file, but ions cannot get through even though they are small enough in principle, because they would first have to lose their hydration spheres, which is too energetically costly.7
How ions become desolvated in nanopores seems also to bear on the performance of devices called supercapacitors, which can rapidly store and discharge large amounts of electrical charge. Unlike ordinary capacitors, where the charge is held on metal plates separated by an insulator, in supercapacitors it is captured in the form of ions absorbed at the surface of porous carbon electrodes with huge surface area. It was predicted that the energy-storage capacity of supercapacitors would decrease as the pores approach nanometre-scale widths, because ions solvated by the bulky ionic liquids used in such devices cannot get inside. But in 2006 a team working in the US and France found the opposite: for some types of porous carbon the capacity increases dramatically for very small pores. The researchers assumed that the ions are somehow stripped bare inside the pores, but they still do not understand quite how that happens.8
It remains to be seen how many problems like this Resolv can resolve. But the program itself aspires to be as much a catalyst as a solution. Havenith says she hopes Resolv will ‘impose long-lasting change’ – that it will stimulate researchers in different fields to share tools and ideas, to recognize that they face some of the same problems and that they are in fact comrades in this new – and old – branch of chemistry.
Philip Ball is a science writer based in London, UK
References
1 Y L A Rezus and H J Bakker, Phys. Rev. Lett., 2007, 99, 148301 (DOI: 10.1103/PhysRevLett.99.148301)
2 J G Davis et al, Nature, 2012, 491, 582 (DOI: 10.1038/nature11570)
3 D Chandler, Nature, 2002, 417, 491 (DOI: 10.1038/417491a)
4 R Zhou et al, Science, 2004, 305, 1605 (DOI: 10.1126/science.1101176)
5 P Liu et al, Nature, 2005, 437, 159 (DOI: 10.1038/nature03926)
6 M Grossman et al, Nat. Struct. Mol. Biol., 2011, 18, 1102 (DOI: 10.1038/nsmb.2120)
7 F Fornasiero et al, Proc. Natl Acad. Sci. USA, 2008, 105, 17250 (DOI: 10.1073/pnas.0710437105)
8 J Chmiola et al, Science, 2006, 313, 1760 (DOI: 10.1126/science.1132195)










No comments yet