Can protein in dinosaur bones survive for millions of years? Rachel Brazil explores the evidence
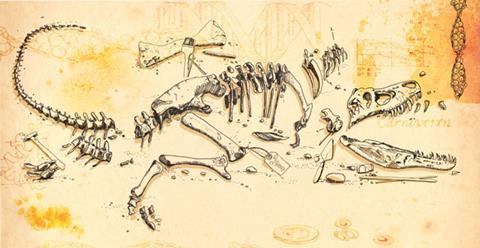
In 2007, Science published protein sequences from a 68-million-year-old dinosaur femur fragment. The bone came from a Tyrannosaurus rex – its name, ‘king of the tyrant lizards’, recognising its one-time status as the largest carnivorous dinosaurs known. The paper, from palaeontologist Mary Schweitzer at North Carolina State University in the US and colleagues reported that amino acid sequences from dinosaur collagen found in the bone closely matched those found in modern chickens. The evolutionary link between birds and dinosaurs was not unexpected, but the results were still met with deep scepticism – could proteins really survive this long?
Although not the first report of ancient protein sequences, Schweitzer’s protein was 100 times older than anything else reported at that time. Within the next 18 months, two rebuttals were published. Critics focused on the quality and statistical analysis of the mass spectrometry data used to identify the protein sequences. Matthew Collins, head of the interdisciplinary biology, chemistry and archaeology group at University of York, UK, was a signatory to one of the rebuttals. ‘Our analysis suggested that collagen would really not survive more than 7 million years at best,’ he says. The feeling was that the evidence was just not strong enough to support a claim this extraordinary.
Alternative explanations for the results were that the proteins came from bacteria or contamination from modern protein. It was also suggested they were really from ostrich samples that were known to have been analysed at co-author John Asara’s mass spectrometry lab at Beth Israel Deaconess Medical Center in Boston, US. Although this took place over a year before the T. rex analysis. Collins says he suspected contamination because the sequences seemed too complete. ‘Old proteins get damaged and destroyed, and there is no evidence of any damage in those peptides – they looked fresh and that would be surprising given their age.’
A stunning discovery
Schweitzer’s involvement in ancient proteomics was serendipitous. She had studied with the renowned US palaeontologist Jack Horner, but her young family made fieldwork difficult so she chose to specialise in lab-based fossil analysis. The T. rex samples were found in Montana by Horner, buried deep in sandstone. Schweitzer says she was stunned at what she saw. ‘I found what looked like blood cells inside the dinosaur bone, it was still very much bone. We discovered that if we demineralised the bone, we had material remaining which also flies in the face of how we thought fossils formed.’ The received wisdom is that organic matter degrades quickly with minerals completely filling the void over time.
She moved on to analysing what was there, using antibodies that can recognise specific proteins and peptides as well as working with Asara to sequence the proteins using liquid chromatography–tandem mass spectrometry (LC–MS/MS), where two mass spectrometers are used in series to improve resolution. To obtain the proteomics data her samples were demineralised using a chelating agent and then the remaining organics chemically extracted before analysis. The amino acid sequences were reconstituted from software-based algorithms and databases of previously sequenced proteins.
Collagen, the fibrous structural protein found in animal connective tissue and bone, makes up 85–90% of the organic material in bone so it is the protein you would most expect to find. But the team also identified other proteins found in bone, including actin, tubulin and histones. ’We do have a haemoglobin sequence, but we only got one sequence, one time, and so I haven’t published it,’ Schweitzer adds.
Standards of evidence in ancient proteomics is a sticky issue, not just because of Schweitzer’s controversial results, but because of the early years of the ancient DNA studies. In the early 1990s, several publications purported to have discovered dinosaur DNA from bones and insects trapped within amber, but these all turned out to be modern DNA contamination. This led to a period of ultra-conservatism and in-fighting, with all results questioned. The field did not recover until the arrival of next generation sequencing, in the last decade.
The ancient proteomics community is now trying hard to avoid repeating earlier mistakes. ‘We don’t want to follow our elder brothers,’ says Enrico Cappellini, a geogenetics researcher at the University of Copenhagen, Denmark. Proteomics researchers are already in a better position than early ancient DNA researchers, due to improvements in mass spectrometry instrumentation. ‘The resolution and sensitivity is so good now that we can see things we never saw before,’ says Collins. Also, contamination is less of a problem with proteomics because it does not involve an amplification step – which is necessary for DNA sequencing. By replicating the DNA sample using PCR (polymerase chain reaction), there is enough to detect, but this also amplifies any contaminants introduced.
Resisting degradation
It is also now clear that protein survives longer than DNA. ‘In a sample it is possible to find protein fragments even when DNA cannot be recorded anymore,’ says Cappellini. Schweitzer’s results aside, the next oldest protein sequence – collagen from a 3.5-million-year-old high arctic camel was reported in 2013 by bioarchaeologist Mike Buckley at University of Manchester, UK. This compares to the oldest sequenced DNA sample – a 700,000-year-old horse found in Canada’s Yukon territory. And it’s not just collagen being found – in 2012, Cappellini identified 126 distinct protein sequences from a 43,000-year-old woolly mammoth bone.
I started looking at the chemistry behind these degradation-resistant proteins – it might essentially be the same process we use to fix tissues
It’s still not clear how bone proteins are resisting degradation to survive millions of years. With collagen, Buckley believes there are two factors explaining its strength – the protective encapsulating effect of the inorganic hydroxyapatite in bone, but also the structure of collagen itself which is made up of a triple helix of polypeptides. ‘It has this hierarchical structure with these triple helices, themselves crosslinked with other helices, forming large arrays of fibrils and fibres making a strong rope-like structure,’ he explains.
Schweitzer has another theory related to the chemical activity of iron found in blood. She has identified goethite-like (a-FeOOH) nanoparticles in the fossil soft tissue, which she believes may have been derived from haemoglobin. She says an idea came to her when listening to a lecture on the possible link between Alzheimer’s disease and high iron levels in the brain. ‘I started looking at the chemistry behind that.’ She wondered if reactive iron released from haemoglobin would generate free radicals leading to bonds forming between proteins. ‘Essentially the same process occurs when we use formaldehyde to fix tissues – you just crosslink everything so that nothing can get at it to degrade it,’ she says. Formaldehyde crosslinks primary amino groups in proteins with other nearby nitrogen atoms through a carbon link. Schweitzer has tested this idea using modern blood vessels in a haemoglobin solution. ‘It’s been about six years now and they are still very happily sitting in the lab at room temperature.’ The important part is to stop the initial few years of degradation. ‘If you can get them stabilised over that period, what remains can be secondarily stabilised by mineral deposition,’ she adds.
Aiding species identification
In lieu of DNA, archaeologists are using protein sequencing to provide a biological lens further back in time. Buckley was looking for a way to identify and differentiate animal species. Initially, collagen seemed like an unlikely solution, because it has a consistent repeating amino-acid sequence of glycine, proline and hydroxyproline. But whilst two of collagen’s three helical polypeptide chains are identical, Buckley spotted that the third chain showed much greater variation between species. ‘I started to develop techniques to preferentially isolate some of the fragments from this third chain,’ says Buckley, who used this to create species fingerprints.
He was able to differentiate neolithic bone fragments from goats and sheep excavated in Turkey (a useful distinction for archaeologists studying early societies, as tending sheep needs greater resources). Buckley says the differences in their collagen are small – just two amino acids in one peptide, ‘a sheep and a goat, for example, diverged around 8 million years ago and we rely on one single peptide change – so it’s nowhere near as high resolution as DNA.’
Another long-standing puzzle solved is the taxonomy of South American ungulates – a group of mammals extinct for 10,000 years. In 2015, Collins reported the analysis of samples from the American Museum of Natural History. ‘These things are so weird, because they evolved in isolation – they have features that make them look like a whole range of different animals,’ he explains. He describes them as rodent-like with continually growing teeth, but the size of hippos. When no DNA could be recovered, the York team were brought in to look for proteins. Collagen sequencing confirmed that they were related to the horse family, rather than elephants as had been suggested.

But ancient proteomics has more to offer than species identification, says Cappellini. ‘It allows you to investigate the products of gene expression [proteins] – so whilst genes may be identical, gene expression changes in association with different biological processes.’ This has led to some exciting results from human teeth – or more specifically calcified dental plaque or calculus. Cappellini’s 2014 study of 1000-year-old calculus from a medieval German monastery produced sequences for 43 human proteins. 25 were proteins produced by the immune system to fight invading bacteria, as well as inflammatory proteins that suggest these individuals had active tooth decay. The same team has also sequenced proteins from dental calculus to gain information on diet. Our ancestors were lactose intolerant, but a genetic mutation changed that 4000–5000 years ago and they began to consume raw milk. ‘We are now finding the raw milk protein b-lactoglobulin – that’s a phenomenal marker,’ says Collins. ‘In the past we’ve speculated about the consumption of milk, now we can actually see it in people’s teeth by sequencing the protein.’
More evidence required
Ancient proteomics is now a growing field, but there is still no consensus on the authenticity of the T. rex proteins. In 2009, Schweitzer followed up with a report of proteins from an 80-million-year- old Brachylophosaurus femur. The Brachylophosaurus was a mid-sized dinosaur with a paddle-like plate over the top of its skull. Schweitzer carried out antibody studies and mass spectrometry, now using a clean-lab environment to reduce the risk of contamination as well as higher resolution instrumentation. The data also confirmed the presence of collagen sequences and supported their link to bird collagen.
But that work did not silence her critics. ‘I think a large part of the scientific community is still undecided,’ explains Buckley. He accepts that the likely survival limit for proteins is older than his 3.5-million-year-old high arctic camel. ‘I wouldn’t be overly surprised if it was twice as long, but from a few million years to pushing on 80 million years is a huge jump.’ Collins agrees that this is what current publications suggest, but tantalisingly claims his lab is working on some exciting new ideas as to how proteins may survive longer than this, which he thinks could be game-changing.
Sergio Bertazzo, a chemist and electron microscopy specialist at University College London in the UK, is not a sceptic of Schweitzer’s work. People can always query mass spectrometry results, but when you look at the microscopy images all the evidence for the cell and collagen structures is there, he explains. Bertazzo’s ‘day job’ is imaging diseased blood vessels, and he became involved in ancient bone through a chance meeting with Imperial College London palaeontologist Susannah Maidment. She got hold of some very fragmented Cretaceous (between 145.5 and 65.5 million years ago) dinosaur bone for Bertazzo to image.
When I saw the fibres there with banding, I thought this really looks how the original protein would look
In small regions of the bone, Bertazzo was surprised to clearly see the stripped pattern characteristic of the rope-like collagen fibres he had seen in modern bone. He was convinced he was seeing dinosaur collagen. ‘When I saw the fibres there with banding, I thought this really looks how the original protein would look.’ Energy-dispersive x-ray spectroscopy data confirmed these regions were high in carbon and he decided to take a closer look at the composition using nano-analytical techniques. Bertazzo used surface-sensitive time-of-flight secondary ion mass spectrometry (TOF-SIMS) to reveal areas containing amino-acid fragments typical of collagen fibres. Scanning electron microscopy images also showed areas that looked like red blood cells, and these regions had mass spectrometry signatures that most closely matched the spectra of emu blood.
What Bertazzo is keen to point out is that his fossil samples were not exceptionally well preserved, like those of Schweitzer’s. They were the unclassified odds and ends that the UK’s Natural History Museum were prepared to part with. ‘In micro regions that are for some reason protected, a little of the original organic components are preserved,’ Bertazzo says. ‘We really think that this is common.’
The question then arises – could there still be remaining fragments of dinosaur DNA? Schweitzer has tested for DNA in several ways, including with fluorescent stains such as DAPI (4’,6-diamidino-2-phenylindole), which binds strongly to adenosine and thymine nucleotide-rich regions in double-stranded DNA. The results show fluorescing areas inside the bone cells in patterns comparable to modern cell samples and Schweitzer thinks DNA remnants could still exist. ‘The better question is can we sequence DNA that is older than a million years and then it becomes a technical problem.’ Current next-generation methods cannot sequence DNA fragments of less than 50 base-pairs so it’s unlikely any remaining fragments could be sequenced until technology advances.
In the meantime, analysing ancient proteins is helping archaeologists and palaeontologists pinpoint evidence where it’s sometimes difficult to find. For example, archaeologists are unable to identify the species of 80–90% of bone fragments found on archaeological sites because they are too small. So Buckley is starting to develop a high throughput method for analysing proteins in bone fragments. ‘My lab has analysed over 10,000 samples for a single project,’ he says. This could be particularly useful in looking for ancient human remains which can sometimes be hard to pick out from other animal bones.
Meanwhile, Schweitzer is waiting for more excavated samples to test and it remains to be seen if further evidence or new theories on protein preservation can sway her sceptics. Dinosaurs have been her passion since the age of five, but she thinks there are broader reasons to keep looking for the molecular make-up of long-extinct species. ‘Dinosaurs survived some of the biggest global climate change that any organism has ever seen,’ she says. Understanding more about their variation over time could teach us some general lessons about biology and how it adapts to a changing environment.
Rachel Brazil is a science writer based in London, UK












No comments yet