Simple sugars discriminate between pathogens
Scientists in the UK have developed a new tool to distinguish bacterial strains from each other. This speedy method could help to minimise drug resistance by accurately directing antibiotic treatment at an early stage.
Antimicrobial resistance is a major health concern and its risk is growing evermore due to a lack of both new drugs and rapid point-of-care diagnostic tools to ensure best use of the drugs in hand. ‘Innovative solutions to identifying pathogens can be found – which is really needed as antibiotic resistance spreads – if different fields work together,’ explains Matthew Gibson from the University of Warwick, whose group developed the new diagnostic tool.
Current point-of-care methods to identify bacteria require culturing bacteria to grow them to higher density – a very slow process. New sequencing technologies are faster, but still require hours and special equipment. Gibson’s method makes use of adhesion between bacteria and other cells. Many bacteria bind to cells through protein or carbohydrate structures exposed on the surface of the bacterium called adhesins. Each bacterial strain has a very specific pattern of adhesins and therefore binds with different strength to different sugar-bearing cell surfaces.
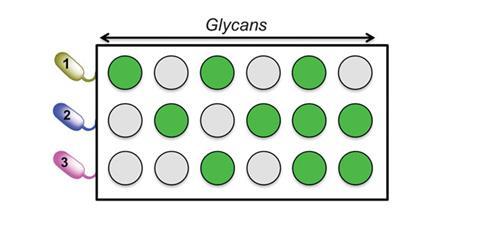
The tool consists of a variety of simple carbohydrates immobilised on a plastic plate, to mimic cell surfaces. The team incubated different strains of bacteria on different plates. They then measured how many bacteria bind to the surface and could determine the relative binding strengths of the different bacteria to these different carbohydrate surfaces. By comparing the data, Gibson’s team obtained a specific ‘barcode’ for each bacterium, describing its binding to those carbohydrate surfaces.
Once a library of barcodes is set up for a variety of bacteria, it is possible to identify any unknown sample of a bacterium as long as it is in the database. ‘[The tool] demonstrates how even simple sugars, when used in combination with ratiometric analyses, can be employed to discriminate between bacterial species. The accessibility of the sugars required makes this an attractive approach for translation’, says Rob Field, an expert on carbohydrate-based sensors from the John Innes Centre in Norwich, UK.
This new analytical method is not only quicker than already existing methods but it is also much cheaper. In the future, Gibson plans to develop the assay into a disposable device to quickly determine bacterial strains, diagnose drug resistance or even monitor known outbreaks.
References
This paper is open access
L Otten, E Fullam and M I Gibson, Mol. BioSyst., 2016, DOI: 10.1039/c5mb00720h











No comments yet