Scientists make 32 nucleotide bricks that can snap together to build huge numbers of three-dimensional shapes
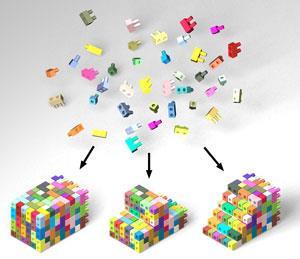
DNA building blocks have been pieced together to form a multitude of Lego-like bricks by researchers at Harvard University and Massachusetts Institute of Technology in the US. The bricks are nanoscopic but can be locked together like children’s Lego blocks and have been used to create more than one hundred distinct shapes with complex surfaces, as well interiors with pores and tunnels.
Chemists have always had the option of turning to natural molecular recognition and self-assembly to help them build novel structures and devices. A wide range of structures is possible using so-called DNA origami for instance. However, while this approach has not been limited to flat, two-dimensional DNA tiles and highly complex structures are possible, the process is not modular. DNA building blocks offer the possibility of constructing much more sophisticated structures and molecular devices.
Peng Yin and colleagues describe their approach to making structures from DNA bricks as both simple and robust. They start with 32 nucleotide strands comprising two head domains and two tail domains each of which is 8 bases in length. They liken the tail-end of each brick to the protruding knobble on a child’s Lego brick that is pushed into the hole of another brick. With an almost limitless range 32 nucleotide bricks available they can construct myriad three-dimensional structures by sticking the bricks together. Protector bricks close off holes and knobbles and prevent additional bricks from attaching themselves to the 3D structure once it is complete.
By treating each 8-base pair as a ‘voxel’ – the volume equivalent of a pixel – in a 10x10x10 three-dimensional array, the team can use a computer to construct a specific shape in silico. The computer then tells them which specific strands present in which bricks from their construction set are needed to build that shape. These are then mixed together, self-assemble and are annealed into the structure in a single step. Using this approach the team has made 100 three-dimensional shapes representing the characters of the English alphabet and the number 0 to 9, as well as arbitrary shapes such as faces and blocks with specific patterns of channels and holes. The team suggests that the approach could be extended to bigger or more sophisticated bricks given adequate computing power for the design process.
DNA origami expert Hendrik Dietz of the Technical University of Munich, Germany, says: ‘The 3D thing is so awesome, I almost got tears in my eyes because of the joy. I love the Lego figures. When looking at this, one cannot help but submit to the power of DNA. Incredibly stunning results.’ However, he adds that it remains to be seen whether these DNA blocks are practical enough to produce high quality shapes useful to nanotechnologists.











No comments yet