Researchers have pinned down the precise position of hydrogen atoms in crystal structures using widely accessible methods
Hydrogen atoms are crucially important for everything from the physical properties of water to the biological function of DNA and proteins. And yet in crystal structures of proteins, and even in small organic molecules, catching sight of them is tricky. Now, researchers have refined conventional techniques so that crystallographers everywhere can build up an accurate picture of where hydrogen atoms are located – at least for small molecules.1
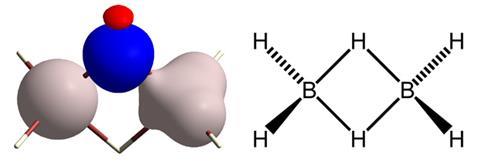
Obtaining a good picture of the position of a molecule’s hydrogen atoms is normally only possible using neutron diffraction, which is only available in very few facilities at an immense cost. Simon Grabowsky from the University of Bremen, Germany, together with colleagues from Poland and Australia, worked with conventional crystallographic datasets, but improved the quantum mechanical modelling required to produce a structure matching the x-ray diffraction data. ‘We use a mathematically more advanced model (Hirshfeld atom refinement) that includes bonding effects, so the experimental data can be reconstructed more successfully and hence the hydrogen atoms be located accurately,’ Grabowsky explains.
Tests applying the method to known crystallographic structures of 81 small molecules show that it determines bond lengths between hydrogens and other atoms as accurately as neutron diffraction. The refinement can be carried out with free software and takes between a few hours and a few days of computation time on an ordinary PC, depending on the size of the molecule.
Louis Farrugia from the University of Glasgow, UK, who was not involved in the new work, notes that some previous attempts to use it with average quality x-ray data were less successful, but he acknowledges that ‘it’s an interesting procedure and with good quality data it certainly gives results close to neutron diffraction standard’. Crystallographer Hideaki Ogata from the Max Planck Institute for Chemical Energy Conversion at Mülheim, Germany, also welcomed the research. ‘The new technique has the advantage that we can see the hydrogens even at resolution as low as 0.8Å,’ he tells Chemistry World.
Hydrogens in proteins offer an even bigger challenge. They often don’t show up at all, although Ogata and colleagues recently revealed the majority of the hydrogens in a hydrogenase enzyme with data of exceptionally good resolution – in a study motivated by the functional importance of hydrogen in that enzyme.2
Grabowsky cautions that the Hirshfeld method is not suitable for proteins, as disorder remains a problem even in crystals. ‘In Hirshfeld atom refinement we need to calculate a quantum-mechanical wavefunction for the molecule that is to be refined,’ Grabowsky explains. ‘There is no way to calculate wavefunctions of moving particles or overlapping molecules. The wavefunction is always static and always only applies to a single molecule or molecule assembly.’ Consequently, protein crystallographers who want to know exact bond lengths for hydrogen atoms are stuck with neutron diffraction for now.
References
1 M Woinska et al, Sci. Adv., 2016, 2, e1600192 (DOI: 10.1126/sciadv.1600192)
2 H Ogata, K Nishikawa and W Lubitz, Nature, 2015, 520, 571 (DOI: 10.1038/nature14110)









No comments yet