New twist on x-ray crystallography could simply reveal functional regions of proteins

A new imaging technique that takes x-ray snapshots of proteins as they twitch in electric fields could let scientists simply unravel which parts of protein give them their function. The US team tested the technique on a common human protein, discovering new information about it. The research may prove useful for guiding rational protein and drug design.
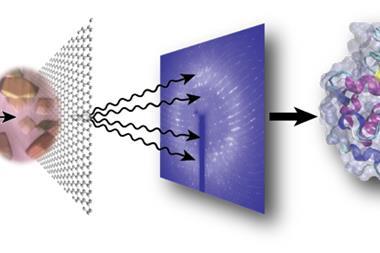
X-ray crystallography has been the gold standard for examining biomolecules since the days of Rosalind Franklin, whose work provided crucial insights into the structure of DNA, and Dorothy Hodgkin, who solved the structure of insulin. The technique works by passing x-rays through a crystal of the material and using the resulting diffraction pattern to infer its molecular structure. However, it is often unclear how the structure of a protein enables it to perform its job. Some techniques can track proteins in motion, like time-resolved crystallography that looks at how the diffraction pattern evolves when the molecule is excited. For proteins containing light-absorbent groups called chromophores, this is often achieved by hitting the molecules with a light pulse to kick-start a process and then using rapid x-ray pulses to follow the motion. However, this deposits biologically implausible amounts of energy in one part of the molecule. It is also useless for proteins that contain no chromophores.
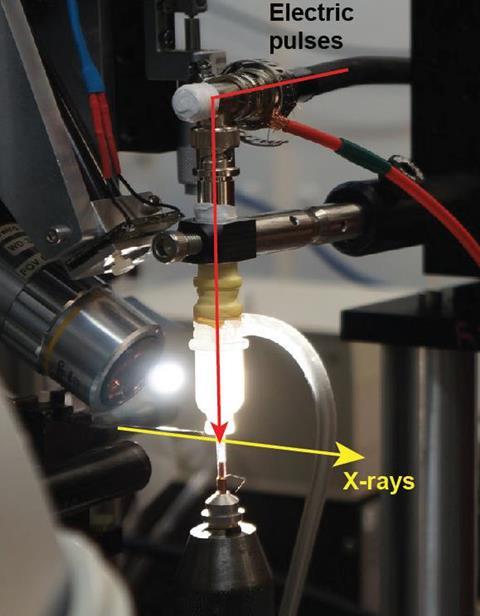
Protein mechanics such as ligand binding are – like all chemical reactions – controlled by electromagnetic forces. So, to study their motion under more physiologically realistic conditions, Rama Ranganathan of UT Southwestern Medical Center and colleagues developed a new technique by sandwiching protein crystals between two glass capillaries containing wires. They then applied electric fields across the crystals that were equivalent to those experienced by tissues. The researchers applied microsecond-scale electric pulses while using x-rays from a synchrotron facility to record diffraction patterns at the instant before the electric pulse hit, as well as at 50, 100 and 200 nanoseconds afterwards. ‘The method is applicable to any protein and the forces that we’re putting in are much more like the small forces that drive protein conformational changes in natural settings,’ says Ranganathan.
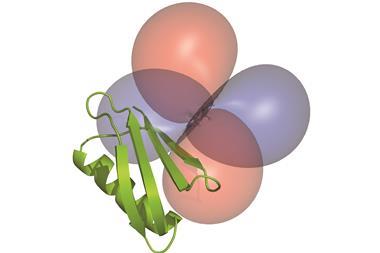
The researchers studied part of a human ubiquitin ligase protein, which binds to peptide ligands. They found that the parts of this protein that moved most in response to the electric field were those known to perform an evolutionarily conserved role binding ligands across a variety of distantly-related domains in proteins from many different species. This suggests that perturbing the structure of a protein and watching how it moves could show researchers the functionally important motions it has evolved to be able to perform. ‘That remains a deep hypothesis that needs to be tested much further,’ says Ranganathan. ‘But I think the finding that the electric-field induced motions are correlated to the ligand binding effect tells us that we’re on the right track.’
‘The technology of what they did and what they found is really interesting,’ says structural biologist Elizabeth Getzoff of the Scripps Research Institute, US, who was not involved in the work. She describes the association between observed motions and ligand binding as ‘exciting’. She cautions, however, that the researchers rely on specific features of the protein crystal that are not universal. ‘It’s not clear to me how generalisable this will be to other systems,’ she adds.
References
D R Hekstra et al, Nature, 2016, DOI: 10.1038/nature20571









1 Reader's comment