An artificial protein that self-assembles around and protects DNA could be ideal for gene therapy, nanomachines and synthetic biology
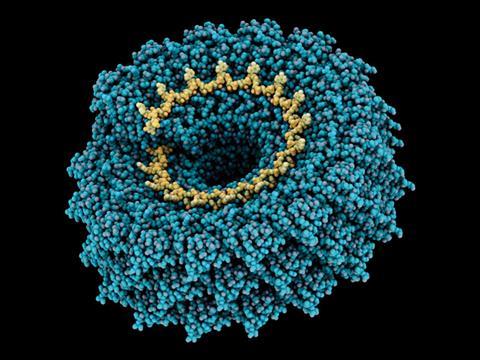
Dutch scientists have built a simple model of viruses’ protective coats in an attempt to create viral mimics that could fight diseases, as opposed to causing them. Rather than copying natural proteins, Renko de Vries from Wageningen University and his team designed and built a three-part protein from scratch that self-assembles around DNA.
‘The protein is exceedingly simple in its primary and secondary structure, yet captures the essence of self-assembly for the tobacco mosaic virus,’ de Vries tells Chemistry World. This knowledge could enable superior vehicles for getting DNA and RNA into cells, for example for gene therapy, and templates for improved DNA machines. ‘You could probably do the same with supramolecular chemistry,’ de Vries adds, ‘but the protein approach has the beauty that you can expand in the direction of synthetic biology.’
The ‘no-frills’ coat sprung from de Vries’ discussions with Paul van der Schoot’s Technical University of Eindhoven team, who had developed a theoretical model of tobacco mosaic virus self-assembly. ‘We established the crucial mechanisms and then started designing these molecules,’ de Vries explains.
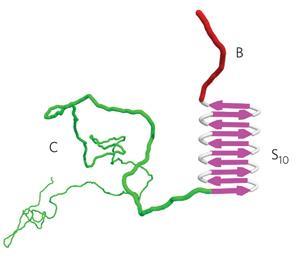
The protein’s first segment, which bound to the DNA to be encapsulated, simply comprised 12 lysine amino acid building blocks. The second was a ‘silk-like’ protein sequence, containing repeat units of mostly alanine and glycine amino acids, that can form stiff filaments. Varying the number of repeat silk-like units allowed the chemists to dictate cooperation between segments during coat assembly. The third segment was a random 400 residue sequence with many prolines and other hydrophilic, uncharged amino acids that stopped the rod-shaped ‘virus-like particles’ (VLPs) clumping together.
Guaranteed well-dressed
‘We found that the self-assembly was really quite spectacular,’ de Vries recalls. ‘If you have one protein sticking to the nucleic acid template, that accelerates binding of further proteins. That ensures that you always have at least a couple of templates perfectly coated, even if you do not have enough protein. For the 2500 base pair linear DNA we used, about 400 copies of the artificial virus protein are needed to make the complete coat.’
Out of five different silk-like segment lengths the team tried, only the two longest ones led to fully cooperative coat self-assembly. These VLPs compacted their central DNA most and protected it from enzyme attack for longer. However, all of the different silk-like segment lengths produced VLPs that could transfect DNA into cells with similar efficiency.
De Vries emphasises that despite improved self-assembly having little impact on transfection, this doesn’t undermine his team’s findings, as they weren’t after a good DNA delivery vehicle. ‘It was our aim to mimic the self-assembly process,’ he says. ‘If you have a very precise scaffold, then that offers opportunities to start mimicking other aspects of viruses, like how they get into and are transported within cells.’ However, his team does want better control of how the DNA triggers self-assembly. ‘Right now nucleation happens at the end of the DNA template, but that’s more-or-less coincidental.’
Yongju Kim, who studies self-assembling biological molecules at Jilin University in China, calls the work ‘amazing’. ‘It really seems to be like natural viruses, especially if these artificial viruses can show some examples able to recognise specific DNA sequences,’ he says. ‘It will be a stepping stone and provide many insights for further artificial virus study.’
Sierin Lim, a protein engineer at Nanyang Technological University in Singapore, is also impressed by the work. She says that it’s exciting that the team has brought rational design to functions like DNA protection and transfection. ‘There will be many potential applications waiting to be explored using this platform,’ says Lim.











No comments yet